Hello....We are just starting to transfer our microsatellite projects over to NGS and were planning on using this program to genotype our samples. We have a couple of ongoing projects were we add to the datasets every year so we are transferring our legacy loci over to our MiSeq. I was trying to test out the program using the test data on our computers to make sure I can get it to work but have been unable to do so far. I am using the primer-input txt and the ErrorRate_FASTQ.gz_files as my data set folder.
I am using a Windows 10 with 64-bit OS and have also tried the program on an Mac but got the same results. I followed the instructions in the manual on how to install...just decompressed the zip folder.
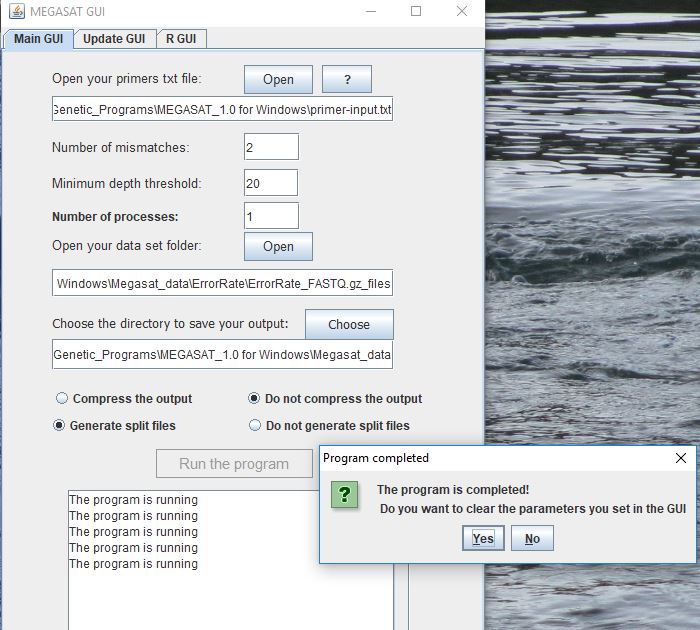
This is a screenshot of the GUI window. Once I hit Run the program, immediately another window appears saying the program is complete but there is nothing written in the output directory. I had a colleague try to run it on his windows machine but he got an error saying he needed the Pearl: ParallelForkManager installed.
If you have any ideas on what I am doing wrong or if there is something else I need installed on my computer that would be greatly appreciated.
thanks
Rob
Hello....We are just starting to transfer our microsatellite projects over to NGS and were planning on using this program to genotype our samples. We have a couple of ongoing projects were we add to the datasets every year so we are transferring our legacy loci over to our MiSeq. I was trying to test out the program using the test data on our computers to make sure I can get it to work but have been unable to do so far. I am using the primer-input txt and the ErrorRate_FASTQ.gz_files as my data set folder.
I am using a Windows 10 with 64-bit OS and have also tried the program on an Mac but got the same results. I followed the instructions in the manual on how to install...just decompressed the zip folder.
This is a screenshot of the GUI window. Once I hit Run the program, immediately another window appears saying the program is complete but there is nothing written in the output directory. I had a colleague try to run it on his windows machine but he got an error saying he needed the Pearl: ParallelForkManager installed.
If you have any ideas on what I am doing wrong or if there is something else I need installed on my computer that would be greatly appreciated.
thanks
Rob