This article comes from Din, N.U., Yu, J. Training a deep learning model for single-cell segmentation without manual annotation. Sci Rep 11, 23995 (2021). https://doi.org/10.1038/s41598-021-03299-4
Here is only a jupyternotebook to present the result from authors' work by using GPU.
First of all we need to download the specific packages/git clone we need in this task:
!pip install pymaxflow
!git clone https://github.com/jiyuuchc/cellcutter.gitThen we need to load the cell image data(note: for this project's data you can also choose'a2data''a3data' for the test):
# Load some image data
data = np.load('cellcutter/data/a1data.npz')
train_data = data['data']
input_img = train_data[...,0]
nucleus_img = train_data[..., 2]
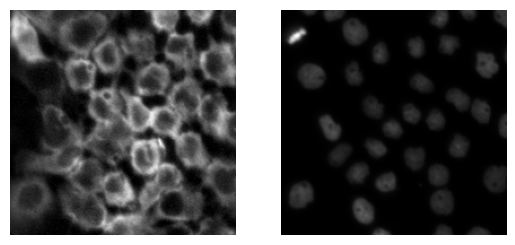
# Check the images
fig, ax = plt.subplots(1,2)
ax[0].imshow(input_img[100:300,200:400])
ax[0].axis('off')
ax[1].imshow(nucleus_img[100:300,200:400])
ax[1].axis('off')The next step is to define the area of anlysis by graphcut:
## Defined the area of analysis
mask = cellcutter.utils.graph_cut(input_img, prior = 0.985, max_weight=10, sigma = 0.03)
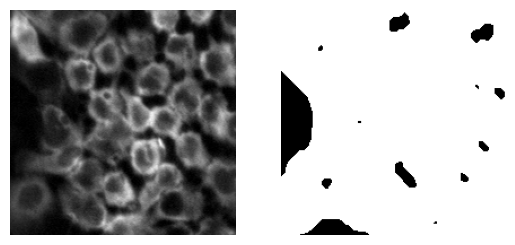
#check results
fig, ax = plt.subplots(1,2)
ax[0].imshow(input_img[100:300,200:400])
ax[0].axis('off')
ax[1].imshow(mask[100:300,200:400])
ax[1].axis('off')-
The function
graph_cuttakes an input image and returns a binary mask that separates the pixels into two classes: foreground and background. -
The
priorparameter is a coefficient that controls the a priori probability of a pixel being in the foreground. -
The
max_weightparameter controls the maximum edge weight between adjacent pixels in the graph. In graph cut algorithms, the image is represented as a graph, where nodes correspond to pixels and edges correspond to the similarity between adjacent pixels. Themax_weightparameter sets an upper bound on the edge weights, which can help to prevent the algorithm from over-segmenting the image. -
The
sigmaparameter is a smoothing parameter that controls the strength of the smoothing applied to the input image before graph construction. It determines the size of the Gaussian blur kernel used to smooth the image. A largersigmavalue will result in a smoother image, while a smaller value will preserve more of the image's detail.
Now we start to train our model:
# CNN segmentation with epochs of 30
dataset = cellcutter.Dataset(input_img, markers, mask_img = ~mask) # actually need the inverse of the mask
start = time.time()
model = cellcutter.UNet4(bn=True)
cellcutter.train_self_supervised(dataset, model, n_epochs = 30)
print('Elapsed time: %f'%(time.time() - start))
# Check the segmentation results
from skimage.color import label2rgb
label = cellcutter.utils.draw_label(dataset, model, np.zeros_like(input_img, dtype=int))
rgb = label2rgb(label, bg_label = 0)
border = cellcutter.utils.draw_border(dataset, model, np.zeros_like(input_img, dtype=int))The training steps will be shown like below:
We can see the loss function will be negative by definition.
Then we want to show our output figure:
# Check the segmentation results
from skimage.color import label2rgb
label = cellcutter.utils.draw_label(dataset, model, np.zeros_like(input_img, dtype=int))
rgb = label2rgb(label, bg_label = 0)
border = cellcutter.utils.draw_border(dataset, model, np.zeros_like(input_img, dtype=int))
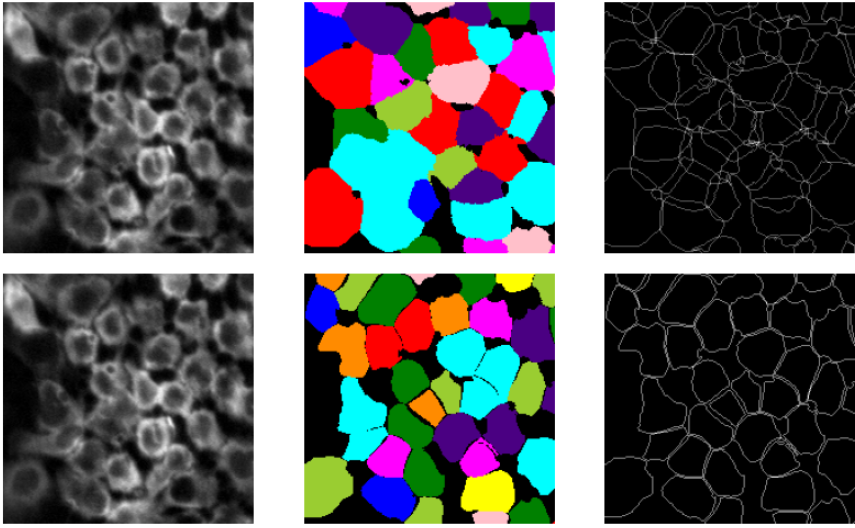
fig, ax = plt.subplots(1,3,figsize = (10,7))
ax[0].imshow(input_img[100:300,200:400])
ax[0].axis('off')
ax[1].imshow(rgb[100:300,200:400])
ax[1].axis('off')
ax[2].imshow(border[100:300,200:400])
ax[2].axis('off')
plt.show()Here we list results of epochs = 10 and epochs = 30 :
We can see the masks and segmentation result get clearer with the increase of epochs.