NEWS 1/9/26: 🎉 the FRD paper has been accepted to Medical Image Analysis 🎉!
FRD (Fréchet Radiomic Distance): A Metric Designed for Medical Image Distribution Comparison in the Age of Deep Learning
By Nicholas Konz*, Richard Osuala*, (* = equal contribution), Preeti Verma, Yuwen Chen, Hanxue Gu, Haoyu Dong, Yaqian Chen, Andrew Marshall, Lidia Garrucho, Kaisar Kushibar, Daniel M. Lang, Gene S. Kim, Lars J. Grimm, John M. Lewin, James S. Duncan, Julia A. Schnabel, Oliver Diaz, Karim Lekadir and Maciej A. Mazurowski.
Project Website · Paper (Medical Image Analysis) · arXiv · Evaluation Framework · Documentation · API
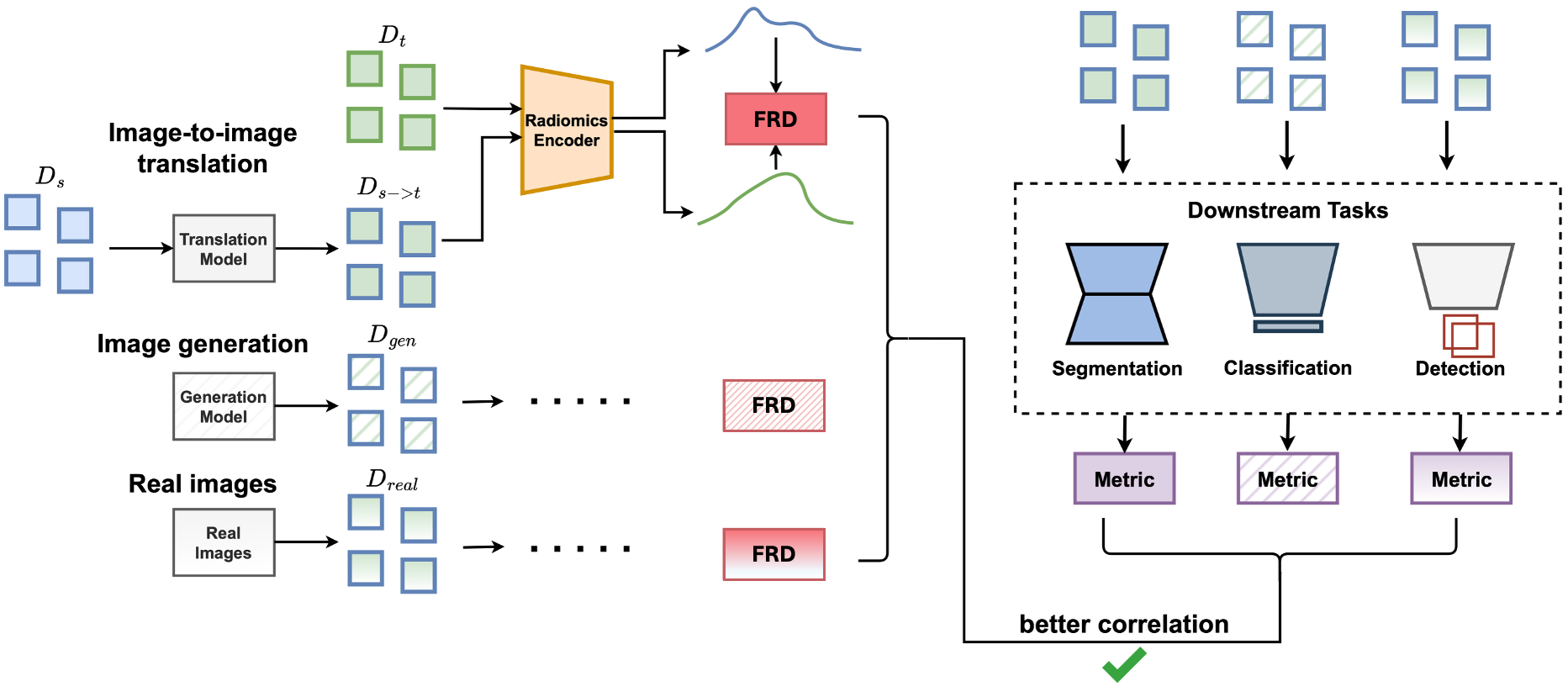
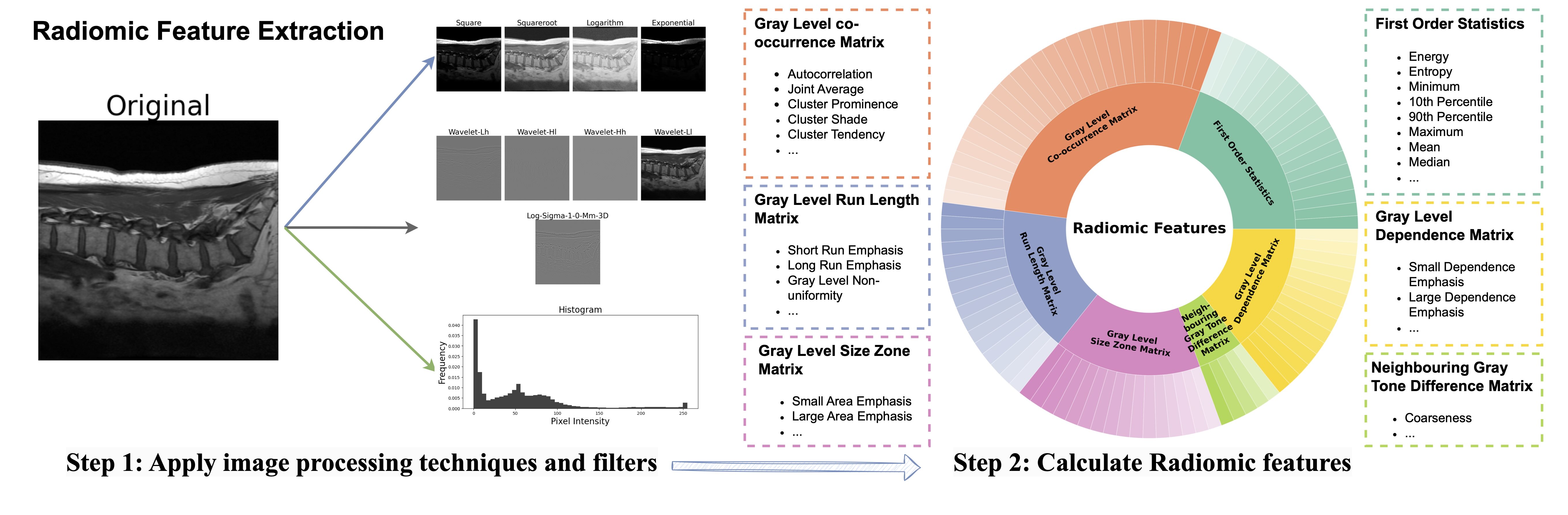
FRD measures the similarity of radiomic image features between two datasets by computing the Fréchet distance between Gaussians fitted to the extracted and normalized features. The lower the FRD, the more similar the two datasets.
FRD supports both 2D (PNG, JPG, TIFF, BMP) and 3D (NIfTI .nii.gz) radiological images.
FRD uses standardised radiomic features rather than pretrained deep features (as in FID, KID, CMMD). We show in our paper that this yields:
- Better alignment with downstream task performance (e.g. segmentation).
- Improved stability and computational efficiency for small-to-moderately-sized datasets.
- Improved interpretability, because radiomic features are clearly defined and widely used in medical imaging.
| FRDv0 (Osuala et al., 2024) | FRDv1 (Konz, Osuala et al., 2026) — default | |
|---|---|---|
| Features | ~94 (Original only) | ~464 (Original + LoG + Wavelet) |
| Normalization | min-max, joint | z-score, D1-referenced |
| Output | raw Fréchet distance | log-transformed Fréchet distance |
| Feature classes | firstorder, glcm, glrlm, gldm, glszm, ngtdm, shape, shape2D | firstorder, glcm, glrlm, glszm, ngtdm |
pip install frd-scoreNote:
frd-scorerequires pyradiomics, which must be installed separately from GitHub because the PyPI release is broken for Python ≥ 3.10 (#903):
pip install git+https://github.com/AIM-Harvard/pyradiomics.git@masterconda install -c conda-forge frd-scoreThe conda-forge package includes all dependencies (including pyradiomics), so no additional installation steps are needed.
git clone https://github.com/RichardObi/frd-score.git
cd frd-score
pip install git+https://github.com/AIM-Harvard/pyradiomics.git@master
pip install -e ".[dev]"- Python ≥ 3.10
- pyradiomics (installed from GitHub, see above)
- numpy, scipy, Pillow, SimpleITK, opencv-contrib-python-headless
Windows users
Building pyradiomics from source requires a C compiler and CMake. Install Visual Studio Build Tools with the "Desktop development with C++" workload, then retry the pip install.
# Compute FRD between two image folders (default: v1)
python -m frd_score path/to/dataset_A path/to/dataset_B
# Use FRDv0 instead
python -m frd_score path/to/dataset_A path/to/dataset_B --frd_version v0
# With masks
python -m frd_score path/to/dataset_A path/to/dataset_B -m path/to/masks_A path/to/masks_B
# Save precomputed statistics to .npz
python -m frd_score --save_stats path/to/dataset path/to/output.npz
# Re-use .npz file
python -m frd_score path/to/output.npz path/to/dataset_Bfrom frd_score import compute_frd
# Basic usage
frd_value = compute_frd(["path/to/dataset_A", "path/to/dataset_B"])
# With masks and options
frd_value = compute_frd(
["path/to/dataset_A", "path/to/dataset_B"],
paths_masks=["path/to/masks_A", "path/to/masks_B"],
frd_version="v1",
verbose=True,
)
# From file lists
frd_value = compute_frd([
["img1.png", "img2.png", "img3.png"],
["img4.png", "img5.png", "img6.png"],
])python -m frd_score path1 path2 [OPTIONS]
| Flag | Description | Default |
|---|---|---|
--frd_version |
v0 or v1 |
v1 |
-m, --paths_masks |
Two mask folder paths | None |
-f, --feature_groups |
Feature classes to extract (e.g. firstorder glcm) |
version default |
-I, --image_types |
Image filter types (Original, LoG, Wavelet) |
version default |
-r, --resize_size |
Resize images to N×N or W×H | None |
-R, --norm_range |
Normalization range [min max] |
version default |
-T, --norm_type |
minmax or zscore |
version default |
--norm_ref |
Normalization reference: joint, d1, independent |
version default |
-v, --verbose |
Verbose logging | off |
-w, --num_workers |
CPU workers for multiprocessing | auto |
-s, --save_stats |
Save statistics to .npz |
off |
-F, --save_features |
Save features to CSV | off |
--use_paper_log |
Use paper Eq. 3 log transform: log(√d²) |
off |
--means_only |
Mean-only Fréchet distance (no covariance) | off |
--log_sigma |
LoG sigma values | [2.0, 3.0, 4.0, 5.0] |
--bin_width |
PyRadiomics bin width | 5 |
--normalize_scale |
PyRadiomics normalize scale | 100 |
--voxel_array_shift |
PyRadiomics voxel array shift | 300 |
--config_path |
Custom PyRadiomics YAML config | None |
--exclude_features |
Post-extraction exclusion: textural, wavelet, firstorder, shape |
None |
--match_sample_count |
Subsample larger dataset to match smaller | off |
--interpret |
Run interpretability analysis | off |
--interpret_dir |
Output dir for interpretation plots | outputs/interpretability_visualizations |
# Image-level OOD detection
python -m frd_score ood path/to/reference path/to/test
# Dataset-level nFRD
python -m frd_score ood path/to/reference path/to/test --detection_type dataset| Flag | Description | Default |
|---|---|---|
--detection_type |
image or dataset |
image |
--val_frac |
Fraction of reference held out for threshold | 0.1 |
--use_val_set |
Enable hold-out validation split | off |
--id_dist_assumption |
gaussian, t, or counting |
gaussian |
--output_dir |
Directory for OOD CSV output | outputs/ood_predictions |
--seed |
Random seed for reproducibility | None |
All shared extraction flags (--frd_version, -f, -I, --norm_ref, etc.) are also available in the ood subcommand.
Main entry point. See API docs for the full signature and parameter descriptions.
Compute and save feature statistics to a .npz file for later re-use. Accepts the same parameters as compute_frd().
Run interpretability analysis on extracted features. Produces t-SNE plots and per-feature difference rankings. Requires matplotlib and scikit-learn.
Out-of-distribution detection using normalized radiomics features. Supports per-image scoring (detection_type="image") and dataset-level nFRD (detection_type="dataset").
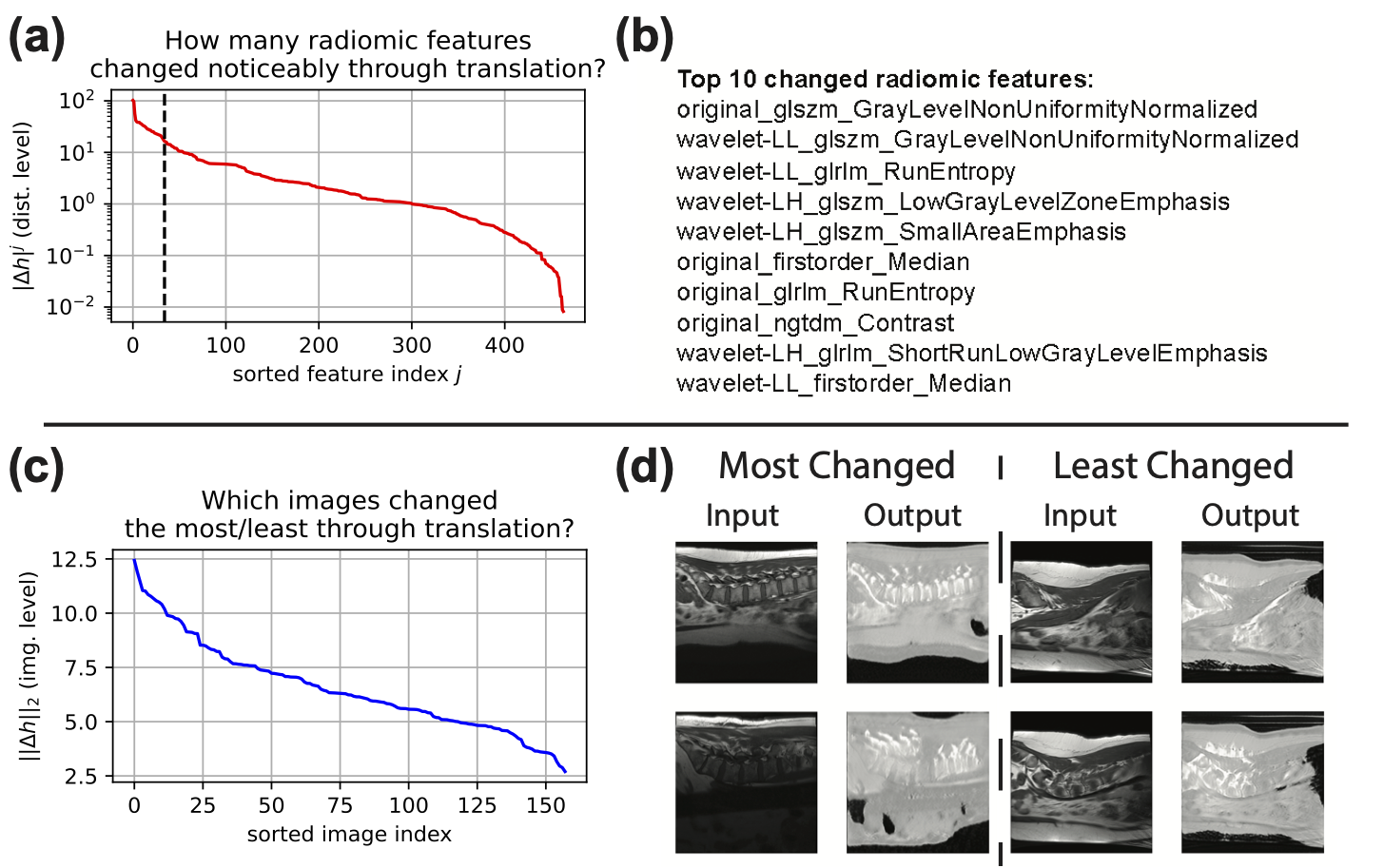
FRD enables interpretable comparison of image sets. Use the --interpret flag or call interpret_frd() to:
- Rank the most-changed radiomic features between two distributions
- Visualise feature distributions via t-SNE
- Identify which images changed the most (for paired datasets)
python -m frd_score path/to/dataset_A path/to/dataset_B --interpretFRD can detect whether newly acquired medical images come from the same domain as a reference set — useful for flagging potential distribution shifts (e.g. different scanners, protocols).
# Per-image OOD scores and p-values
python -m frd_score ood path/to/reference path/to/test_images
# Dataset-level OOD score (nFRD)
python -m frd_score ood path/to/reference path/to/test_images --detection_type datasetResults are saved to outputs/ood_predictions/ood_predictions.csv with columns: filename, ood_score, ood_prediction, p_value.
If you use this library in your research, please cite:
@article{konz2026frd,
title = {Fr\'{e}chet Radiomic Distance (FRD): A Versatile Metric for
Comparing Medical Imaging Datasets},
author = {Konz, Nicholas and Osuala, Richard and Verma, Preeti and
Chen, Yuwen and Gu, Hanxue and Dong, Haoyu and Chen, Yaqian
and Marshall, Andrew and Garrucho, Lidia and Kushibar, Kaisar
and Lang, Daniel M. and Kim, Gene S. and Grimm, Lars J. and
Lewin, John M. and Duncan, James S. and Schnabel, Julia A. and
Diaz, Oliver and Lekadir, Karim and Mazurowski, Maciej A.},
journal = {Medical Image Analysis},
volume = {110},
pages = {103943},
year = {2026},
publisher = {Elsevier},
doi = {10.1016/j.media.2026.103943},
url = {https://www.sciencedirect.com/science/article/pii/S1361841526000125},
}Earlier FRD work:
@article{osuala2024towards,
title = {Towards Learning Contrast Kinetics with Multi-Condition
Latent Diffusion Models},
author = {Osuala, Richard and Lang, Daniel and Verma, Preeti and
Joshi, Smriti and Tsirikoglou, Apostolia and Skorupko, Grzegorz
and Kushibar, Kaisar and Garrucho, Lidia and Pinaya, Walter HL
and Diaz, Oliver and others},
journal = {arXiv preprint arXiv:2403.13890},
year = {2024},
}- API Documentation — overview, benchmarks, datasets, FAQ
- Change Log — overview, benchmarks, datasets, FAQ
- Project Website — overview, benchmarks, datasets, FAQ
- Journal Article — Medical Image Analysis, Vol. 110 (2026)
- arXiv Preprint
- Evaluation Framework — scripts for OOD detection, translation evaluation, and metric comparison
- API Documentation — full docs hosted on GitHub Pages
- Preeti Verma — implementation of a script of an early frd version.
- Nicholas Konz — FRDv1 and RaD repository
- PyRadiomics — radiomic feature extraction backend
- pytorch-fid — Fréchet distance implementation reference