StatsPAI is the agent-native Python package for causal inference and applied econometrics. One import, 800+ functions, covering the complete empirical research workflow — from classical econometrics to cutting-edge ML/AI causal methods to publication-ready tables in Word, Excel, and LaTeX.
Designed for AI agents: every function returns structured result objects with self-describing schemas (list_functions(), describe_function(), function_schema()), making StatsPAI the first econometrics toolkit purpose-built for LLM-driven research workflows — while remaining fully ergonomic for human researchers.
It brings R's Causal Inference Task View (fixest, did, rdrobust, gsynth, DoubleML, MatchIt, CausalImpact, ...) and Stata's core econometrics commands into a single, consistent Python API.
pip install statspai, then run any of the four canonical causal-inference exercises below. StatsPAI ships the classic teaching datasets bundled under sp.datasets — Callaway–Sant'Anna mpdta, Card (1995) returns-to-schooling, Abadie–Diamond–Hainmueller California Prop 99, Lee (2008) Senate RD, LaLonde / NSW–DW, Angrist–Krueger (1991) QOB, Basque terrorism, German reunification — so every snippet runs offline with no data wrangling.
import statspai as sp
sp.datasets.list_datasets() # name / design / n_obs / paper / expected_mainMinimum-wage effect on teen employment (the canonical example used in R's did package).
import statspai as sp
df = sp.datasets.mpdta()
cs = sp.callaway_santanna(data=df, y='lemp', t='year',
i='countyreal', g='first_treat')
print(sp.aggte(cs, type='simple').summary())
# Simple ATT ≈ -0.033, bootstrap SE ≈ 0.004, p < 0.001Instrument endogenous educ with proximity to a 4-year college (nearc4).
import statspai as sp
df = sp.datasets.card_1995()
iv = sp.ivreg('lwage ~ (educ ~ nearc4) + exper + expersq + black + south + smsa',
data=df)
print(iv.summary())
# educ coefficient ≈ 0.142 (SE 0.019); first-stage F ≈ 160; Hausman p ≈ 0.03Sharp RD around the 0-margin cutoff with Calonico–Cattaneo–Titiunik robust bias-corrected inference.
import statspai as sp
df = sp.datasets.lee_2008_senate()
rd = sp.rdrobust(data=df, y='voteshare_next', x='margin', c=0)
print(rd.summary())
# RD estimate ≈ 0.062 (SE 0.024) — incumbent advantage in next-term voteshareAbadie, Diamond & Hainmueller's canonical tobacco-policy evaluation.
import statspai as sp
df = sp.datasets.california_prop99()
sc = sp.synth(data=df, outcome='cigsale', unit='state', time='year',
treated_unit='California', treatment_time=1989)
print(sc.summary())
# Post-1988 ATT ≈ -13.3 packs/capitaBeyond the point estimate, every .summary() above prints inference scaffolding you would otherwise hand-assemble from 3–4 separate R packages:
- DiD —
aggte[simple]carries a uniform critical value (1.96),balanced_e/min_e/max_eevent-time bookkeeping, and 1 000-replication multiplier-bootstrap SEs. - IV — Partial R²(educ) ≈ 0.051 and first-stage F ≈ 160 (comfortably above Stock–Yogo's rule-of-thumb 10), so weak-instrument risk is visible without a second call.
- RD — Conventional (0.073) and Robust bias-corrected (0.062) estimators print side-by-side with effective-sample counts (440 left / 443 right) at the mserd bandwidth.
- Synth — full 12-period treated-vs-counterfactual gap table;
ridge_lambda ≈ 112.9flags that the ASCM (Ben-Michael, Feller & Rothstein 2021) branch is active. Passmethod='adh'to fall back to classical Abadie–Diamond–Hainmueller (ATT ≈ -13.1 on the same panel).
Every result object exposes the same interface — .summary() / .tidy() / .plot() / .to_latex() / .to_docx() / .to_agent_summary() — across all 800+ estimators. For deeper walkthroughs (staggered DiD, weak-IV diagnostics, RD bandwidth choice, 20 synth methods, DML, matching, spatial, ...) see docs/guides/.
StatsPAI's focus is causal inference — and on this axis we aim to be the most complete single package in any language. "Stata" = official + major SSC packages. "R" = CRAN. "sm+lm" = statsmodels + linearmodels.
| Method family | Stata | R | sm+lm | DoubleML | StatsPAI |
|---|---|---|---|---|---|
| DiD — staggered (CS/SA/BJS/dCdH/Gardner/Wooldridge ET) + event-study + honest CIs | ✅ | ❌ | ❌ | 🏆 | |
| IV — classical (2SLS/LIML/GMM) + modern (Kernel IV / Deep IV / KAN-DeepIV) | ✅ classical only | ✅ classical only | 🏆 | ||
| RD — CCT + 2D / boundary + multi-cutoff + honest CIs + ML-CATE (18+ estimators) | ✅ (rdrobust) |
❌ | ❌ | 🏆 | |
| Synthetic Control — ADH / ASCM / gsynth / BSTS / Bayesian / PenSCM / FDID (20 methods) | ❌ | ❌ | 🏆 | ||
| Double / Debiased ML | ❌ | ✅ | ❌ | ✅ | ✅ |
| Meta-Learners (S/T/X/R/DR) + Causal Forest / GRF | ❌ | ✅ | ❌ | ❌ | ✅ |
| TMLE / HAL-TMLE | ❌ | ✅ | ❌ | ❌ | ✅ |
| Neural causal (TARNet / CFRNet / DragonNet) | ❌ | ❌ | ❌ | ❌ | 🏆 |
| Causal discovery (NOTEARS / PC / LiNGAM / GES) | ❌ | ❌ | ❌ | 🏆 | |
| Proximal CI (fortified / bidirectional / MTP / DNC) | ❌ | ❌ | ❌ | 🏆 | |
| QTE / distributional TE / CiC / dist-IV | ❌ | ❌ | ✅ | ||
| Mendelian randomization (IVW/Egger/median/mode/PRESSO/MVMR/BMA) | ❌ | ✅ | ❌ | ❌ | ✅ |
| Conformal causal inference | ❌ | ❌ | ❌ | ❌ | 🏆 |
| Bayesian Causal Forest (BCF / ordinal / factor-exposure) | ❌ | ❌ | ❌ | ✅ | |
| Spatial econometrics (weights → ESDA → ML/GMM → GWR/MGWR → panel) | ❌ | ❌ | ❌ | 🏆 |
Legend: 🏆 most complete across ecosystems · ✅ full coverage ·
StatsPAI at a glance: 889 registered functions · 78 modules · 188,244 LOC (core) + 42,768 LOC (tests). For the full coverage matrix (23 method families), per-module breakdown, and cross-ecosystem line-count comparison — see docs/stats.md.
🎉 NEW in v1.6.0 — P1 Agent-Native × Frontier (LLM-DAG + sp.paper() + Causal-Text + MR frontier + long-panel DML + agent-native infrastructure)
StatsPAI 1.6.0 is a pure-additive minor release pushing two competitive axes simultaneously: agent-native adoption (closed-loop LLM-DAG, end-to-end publication pipeline, 30 populated agent cards, typed exception taxonomy with recovery hints, auto-generated ## For Agents blocks in every flagship guide) and the methodological frontier (five post-2020 Mendelian-randomization estimators, long-panel Double-ML, constrained PC discovery, two causal_text MVPs).
| Area | v1.6 Highlights |
|---|---|
| Agent-native pipeline | sp.paper(data, question, ...) — orchestrator on top of sp.causal() that parses a natural-language question, runs diagnose → recommend → estimate → robustness, and assembles a 7-section PaperDraft (Question / Data / Identification / Estimator / Results / Robustness / References) with .to_markdown() / .to_tex() / .to_docx() / .write(path). Per-section failure isolation: a failed estimator yields a "Pipeline notes" section rather than crashing the draft. Family guide: docs/guides/paper_pipeline.md. |
| LLM × DAG (closed-loop) | sp.llm_dag_constrained — iterate propose → constrained PC → CI-test validate → demote until convergence. Every kept edge carries llm_score + ci_pvalue + source ∈ {required, forbidden, demoted, ci-test}. result.to_dag() round-trips into statspai.dag.DAG. sp.llm_dag_validate audits any declared DAG edge-by-edge for spuriousness. sp.pc_algorithm(forbidden=, required=) injects background knowledge into PC (default None preserves prior contract). Family guide: docs/guides/llm_dag_family.md. |
| Causal × Text (experimental) | sp.text_treatment_effect — Veitch-Wang-Blei (2020 UAI) text-as-treatment ATE via embedding-projected OLS with HC1 SEs; hash embedder default (deterministic, dependency-free), lazy sbert optional. sp.llm_annotator_correct — Egami-Hinck-Stewart-Wei (2024) Hausman-style measurement-error correction for binary LLM-derived treatments; raises IdentificationFailure when the LLM has no information. Both subclass CausalResult and ship full agent-card metadata. Family guide: docs/guides/causal_text_family.md. |
| MR frontier (5 new) | sp.mr_lap (Burgess-Davies-Thompson 2016 sample-overlap-corrected IVW). sp.mr_clust (Foley-Mason-Kirk-Burgess 2021 clustered MR via finite Gaussian mixture on Wald ratios, BIC-selected K). sp.grapple (Wang-Zhao-Bowden-Hemani 2021 profile-likelihood MR with joint weak-instrument + balanced-pleiotropy robustness). sp.mr_cml (Xue-Shen-Pan 2021 constrained maximum-likelihood MR with L0-sparse pleiotropy, MR-cML-BIC). sp.mr_raps (Zhao-Wang-Hemani-Bowden-Small 2020 Annals of Statistics robust adjusted profile score with Tukey biweight loss). sp.mr(method='lap' | 'clust' | 'grapple' | 'cml' | 'raps') dispatcher routes all five. 41 new tests in tests/test_mr_frontier.py. |
| Long-panel Double-ML | sp.dml_panel — Semenova-Chernozhukov (2023) long-panel DML: absorbs unit (+ optional time) fixed effects via within-transform, cross-fits ML nuisance learners with unit-split folds (Liang-Zeger compatible), reports cluster-robust SE at the unit level. Empty-covariate fallback reduces to pure FE-OLS. 13 new tests. |
| Typed exception taxonomy | StatsPAIError root + AssumptionViolation / IdentificationFailure / DataInsufficient / ConvergenceFailure / NumericalInstability / MethodIncompatibility, each carrying recovery_hint, machine-readable diagnostics, and a ranked alternative_functions list. Warning counterparts: StatsPAIWarning / ConvergenceWarning / AssumptionWarning plus a rich-payload sp.exceptions.warn() helper. Domain errors subclass ValueError / RuntimeError → existing except blocks keep working unchanged. 13 call-site migrations already shipped (DID, IV, matching, DML-IRM, synth, Bayesian DML). |
| Agent cards + registry | sp.agent_card(name) and sp.agent_cards(category=None) return pre_conditions / assumptions / failure_modes (symptom + exception + remedy + alternative) / alternatives / typical_n_min for 36 flagship functions (regress, iv, did, callaway_santanna, rdrobust, synth, dml, dml_panel, causal_forest, metalearner, match, tmle, bayes_dml, bayes_did, bayes_iv, proximal, mr, qdid, qte, dose_response, spillover, multi_treatment, network_exposure, paper, llm_dag_constrained, llm_dag_validate, text_treatment_effect, llm_annotator_correct, ...). sp.recommend() auto-consumes them: every recommendation now includes agent_card / pre_conditions / failure_modes / alternatives / typical_n_min, with an auto-warning when n_obs < typical_n_min. |
| Result-object agent hooks | CausalResult.violations() / EconometricResults.violations() inspect stored diagnostics (pre-trend p, first-stage F, McCrary, rhat/ESS/divergences, overlap, SMD) and return flagged items with severity / recovery_hint / alternatives. .to_agent_summary() returns a JSON-ready structured payload (point estimate, coefficients, scalar diagnostics, violations, next-steps) alongside the existing prose .summary() and tidy() DataFrame. |
| Docs × agents | 26 auto-rendered ## For Agents blocks across 19 guides (was 0 pre-v1.5.1) via sp.render_agent_block(name) + scripts/sync_agent_blocks.py (CI-enforced via tests/test_agent_blocks_drift.py). Covers DID / IV / RD / Synth / Matching / DML / Meta-learners / TMLE / Causal-Forest / Bayesian / Proximal / MR / QTE / Interference / Causal-Text / LLM-DAG / Paper-pipeline families. |
Every public signature shipped in v1.5.x is byte-for-byte identical in v1.6.0 — this release is purely additive. Existing call sites that catch ValueError continue to catch AssumptionViolation / DataInsufficient / MethodIncompatibility / IdentificationFailure; catching RuntimeError continues to catch ConvergenceFailure / NumericalInstability. New code should prefer the specific subclasses and attach a recovery_hint so agents can act on failures without parsing error strings.
Previously in v1.5.0 — Interference / Conformal / Mendelian family consolidation
StatsPAI 1.5.0 was a minor release bundling three concurrent improvements to the interference, conformal causal inference, and Mendelian Randomization families: full-family documentation guides, unified dispatchers matching the sp.synth / sp.decompose / sp.dml pattern, and a targeted correctness audit that fixed two silent-wrong-numbers issues.
- Family guides (3 new) —
docs/guides/interference_family.md(9 estimators + decision tree + 5 diagnostics),docs/guides/conformal_family.md(all 10 conformal estimators organised around marginal-coverage guarantee),docs/guides/mendelian_family.md(17 MR functions by IV1/IV2/IV3 assumption hierarchy + worked BMI → T2D example). - Unified dispatchers (3 new) —
sp.mr(method=...)(33 aliases),sp.conformal(kind=...)(29 aliases),sp.interference(design=...)(29 aliases). Byte-for-byte identical to direct calls (30 parity tests guarantee this). ⚠️ Correctness —sp.mr_egger: slope inference fixed tot(n−2)(wasstats.norm). Anti-conservative at smalln_snps; numerically invisible forn_snps ≥ ~100.⚠️ Correctness —sp.mr_presso: MC p-value switched from rawmean(null ≥ obs)to standard(k+1)/(B+1)convention (matches R's MR-PRESSO). No longer silently produces-inf.⚠️ Breaking —sp.mrmodule → function dispatcher. Module access preserved atsp.mendelian. See MIGRATION.md.
Previously in v1.4.2 — correctness patches + Proximal / QTE / Causal-RL family guides
StatsPAI 1.4.2 was a patch release with two silent-wrong-numbers fixes and three family guides:
⚠️ correctness fix —sp.dml_model_averaging√n SE scaling bug. The cross-candidate variance aggregator treated the sample-mean influence-function outer product asVar(θ̂_avg)directly, missing a final/ n. Reported SEs were√ntimes too large; on the canonical n=400 DGP the 95% CI width was 4.20 (nominal ≈ 0.21) and empirical coverage was 100%. After the fix, CI width is 0.21 and coverage is ≈ nominal. Regression guard:tests/test_dml_model_averaging.py::test_se_on_correct_scale.⚠️ correctness fix —sp.gardner_didevent-study reference-category contamination. Stage-2 dummy regression pooled never-treated units and treated units outside the event-study horizon into a single baseline, dragging every event-time coefficient toward the mean of that pool. On a synthetic panel with true τ=2 and strict parallel trends, pre-trends came out ≈ -0.30 (should be 0) and post ≈ +1.72 (should be 2.0). Replaced the Stage-2 regression in event-study mode with direct Borusyak-Jaravel-Spiess-style within-(cohort × relative-time) averaging of the imputed gap. After the fix: pre-trends ≈ +0.01, post ≈ +2.02. Non-event-study single-ATT path was already correct and is unchanged.- Family guides —
docs/guides/proximal_family.md(full Proximal Causal Inference walkthrough),docs/guides/qte_family.md(mean → quantile → distribution),docs/guides/causal_rl_family.md(causal RL vs classical CI). - Formally shipped from v1.4.1 cherry-picks —
tests/test_bridge_full.py(10 end-to-end tests forsp.bridge(kind=...)bridging theorems) anddocs/guides/bridging_theorems.md.
Previously in v1.4.1 — v3-frontier Sprint 3: AKM shock-clustered SE, Claude extended thinking, parity + integration suites, 2 new guides
StatsPAI 1.4.1 is an additive follow-up to 1.4.0 that closes the Sprint 3 items:
- AKM shock-clustered SE —
sp.shift_share_political_panel(cluster='shock')computes the panel-extended Adão-Kolesár-Morales (2019) variance estimator recommended by Park-Xu (2026) §4.2 — typically 3× tighter than unit-clustered SEs in settings with 10–100 industries.diagnostics['akm_se']and a human-readablediagnostics['cluster']label surface the result. - Claude extended thinking for Causal MAS —
sp.causal_llm.anthropic_client(thinking_budget=N)opts into the Claude 4.5 / Opus 4.7 extended-thinking API. The reasoning trace is captured onclient.history[-1]['thinking']for auditability but is not returned tocausal_mas. Handles boththinkingandredacted_thinkingcontent blocks. - Parity + integration test suites —
tests/reference_parity/test_assimilation_parity.py(10 checks on the Kalman / particle backends, incl. Kalman↔particle agreement and Student-t contamination robustness) andtests/integration/test_causal_mas_with_fake_llm.py(11 end-to-end MAS tests usingecho_client+ 3 Claude thinking block-splitter tests mocking the Anthropic SDK). - Two new MkDocs guides —
docs/guides/shift_share_political_panel.md(full panel-IV recipe incl. AKM shock-cluster) anddocs/guides/causal_mas.md(multi-agent LLM causal discovery walkthrough).
All v1.4.0 APIs remain stable; the new surface is strictly additive kwargs.
Previously in v1.4.0 — v3-frontier Sprint 2: panel shift-share, real-LLM adapters, particle-filter assimilation, 3 new guides
StatsPAI 1.4.0 is Sprint 2 of the 知识地图 v3 roadmap. Closes the four secondary items flagged at the end of Sprint 1: multi-period Park-Xu political shift-share, real OpenAI / Anthropic LLM adapters for the Causal MAS discovery agent, a particle-filter backend for causal_kalman to handle non-Gaussian priors and nonlinear dynamics, and three new MkDocs guides covering the v3 frontier. 20 unused-import cleanups across Sprint 1 modules. One CI flake (CausalForest ATE parity test) deflaked by seeding the forest explicitly.
| Area | v1.4 Highlights |
|---|---|
| Panel shift-share IV | sp.shift_share_political_panel — Park-Xu (2026) §4.2 multi-period extension: time-varying shares + time-varying shocks, pooled 2SLS with unit / time / two-way FE, per-period event-study table + aggregate Rotemberg top-K. Recovers τ = 0.30 within 0.003 on synthetic 30×4 panels. |
| Real-LLM adapters (Causal MAS) | sp.causal_llm.openai_client — OpenAI SDK ≥ 1.0 (supports Azure / vLLM / Ollama via base_url). sp.causal_llm.anthropic_client — Anthropic Messages API ≥ 0.30, defaults to claude-opus-4-7. sp.causal_llm.echo_client — deterministic scripted-response client for offline tests. Lazy-imported SDKs → zero new runtime deps on the core package. |
| Particle-filter assimilation | sp.assimilation.particle_filter — bootstrap-SIR particle filter with systematic resampling (Gordon-Salmond-Smith 1993; Douc-Cappé 2005). Non-Gaussian priors, heavy-tailed observation noise, nonlinear dynamics via pluggable callbacks. Agrees with exact Kalman to ~0.003 under Gaussian DGPs. sp.assimilative_causal(..., backend='particle') routes the end-to-end wrapper. |
| Documentation (v3 frontier guides) | docs/guides/synth_experimental.md (Abadie-Zhao inverse-SC workflow), docs/guides/harvest_did.md (Borusyak-Hull-Jaravel harvesting DID), docs/guides/assimilative_ci.md (Nature Comms 2026 streaming CI, Kalman + particle backends). Wired into mkdocs.yml nav. |
| v1.3 stable foundation (carried forward) | 11 2025-2026 frontier methods from Sprint 1: synth_experimental_design, rdrobust(..., bootstrap='rbc'), evidence_without_injustice, target_trial.to_paper(fmt='jama'/'bmj'), harvest_did, bcf_ordinal, bcf_factor_exposure, causal_mas, shift_share_political, causal_kalman. All v1.0 capstone surfaces (sp.bridge, sp.fairness, sp.surrogate, sp.epi, sp.longitudinal, sp.question, full MR suite, TARGET checklist) remain intact. |
| Agent-native platform | sp.list_functions() / sp.describe_function() / sp.function_schema() expose OpenAI/Anthropic tool-calling schemas for 874+ registered estimators. 5 new hand-written FunctionSpec entries this release. sp.agent.mcp_server MCP scaffold lets external LLMs call every StatsPAI function via natural-language tool invocation. |
| CI/CD hygiene | tabulate hard-dep from v1.3.0 carried forward. Deflaked test_forest_ate_recovers_average_tau by seeding the forest explicitly (random_state=0, n_estimators=300, larger n). 2 699+ tests passing across all OS × Python matrix entries. |
Previously in v0.9.2 — Decomposition Analysis: 18 first-class decomposition methods across 13 modules (~6,200 LOC, 54 tests), unified under sp.decompose(method=...). Mean (Blinder-Oaxaca/Gelbach/Fairlie/Bauer-Sinning/Yun), distributional (RIF/FFL/DFL/Machado-Mata/Melly/CFM), inequality (Theil/Atkinson/Dagum/Shapley/Lerman-Yitzhaki), demographic (Kitagawa/Das-Gupta), and causal (gap_closing/mediation_decompose/disparity_decompose). Closed-form influence functions for Theil/Atkinson, weighted O(n log n) Dagum Gini, cross-method consistency checks.
Previously in v0.9.1 — Regression Discontinuity: 18+ RD estimators, diagnostics, and inference methods across 14 modules (~10,300 LOC) — now the most feature-complete RD package in any language. Covers CCT sharp/fuzzy/kink, 2D/boundary RD (rd2d), RDIT, multi-cutoff & multi-score, honest CIs (Armstrong-Kolesar), local randomization (rdrandinf/rdwinselect/rdsensitivity), CJM density tests, Rosenbaum bounds, CATE via rdhte + ML variants (rd_forest/rd_boost/rd_lasso), external-validity extrapolation (Angrist-Rokkanen), power (rdpower/rdsampsi), and a one-click sp.rdsummary() dashboard. 97 RD tests pass; rd/_core.py consolidates kernel/WLS/sandwich primitives from 9 files into one 191-line canonical module.
Previously in v0.9.0 — Synthetic Control: 20 SCM estimators + 6 inference strategies + full research workflow, all behind the unified sp.synth(method=...) dispatcher. Seven new estimators in this release: bayesian_synth (Dirichlet MCMC), bsts_synth / causal_impact (Kalman smoother), penscm (Abadie-L'Hour 2021), fdid (Forward DID), cluster_synth, sparse_synth (LASSO), kernel_synth + kernel_ridge_synth. Research workflow: synth_compare() runs all 20 · synth_recommend() auto-selects · synth_power() + synth_mde() first power-analysis tool for SCM · synth_sensitivity() · synth_report(format='latex'). ASCM re-implemented to Ben-Michael et al. (2021) Eq. 3; Bayesian MCMC Jacobian corrected; 9 release-blocker fixes from a 5-agent review; 144 synth tests passing. Canonical datasets: california_tobacco(), german_reunification(), basque_terrorism(). See the synth guide.
Previously in v0.8.0: Spatial Econometrics Full-Stack — 38 new API symbols covering weights, ESDA, ML/GMM regression, GWR/MGWR, and spatial panel. Plus: local projections, GARCH, ARIMA, BVAR, LiNGAM, GES, optimal matching, cardinality matching, RIF decomposition, mediation sensitivity, Cox frailty, AFT survival, rdpower, survey calibration. 60+ new functions across 10 domains.
Built by the team behind CoPaper.AI · Stanford REAP Program
| Pain point | Stata | R | StatsPAI |
|---|---|---|---|
| Scattered packages | One environment, but $695+/yr license | 20+ packages with incompatible APIs | One import, unified API |
| Publication tables | outreg2 (limited formats) |
modelsummary (best-in-class) |
Word + Excel + LaTeX + HTML in every function |
| Robustness checks | Manual re-runs | Manual re-runs | spec_curve() + robustness_report() — one call |
| Heterogeneity analysis | Manual subgroup splits + forest plots | Manual lapply + ggplot |
subgroup_analysis() with Wald test |
| Modern ML causal | Limited (no DML, no causal forest) | Fragmented (DoubleML, grf, SuperLearner separate) | DML, Causal Forest, Meta-Learners, TMLE, DeepIV |
| Neural causal models | None | None | TARNet, CFRNet, DragonNet |
| Causal discovery | None | pcalg (complex API) |
notears(), pc_algorithm(), lingam(), ges() |
| Spatial econometrics | None | 5 packages (spdep+spatialreg+sphet+splm+GWmodel) | 38 functions: weights→ESDA→ML/GMM→GWR/MGWR→panel |
| Policy learning | None | policytree (standalone) |
policy_tree() + policy_value() |
| Result objects | Inconsistent across commands | Inconsistent across packages | Unified CausalResult with .summary(), .plot(), .to_latex(), .cite() |
| Interactive plot editing | Graph Editor (no code export) | None | sp.interactive() — GUI editing with auto-generated code |
StatsPAI is not a wrapper for R. We independently re-implement every algorithm from the original papers (with citations exposed via .cite()), and for a few mature engines (pyfixest, rdrobust) we use explicit, transparent bindings. What makes StatsPAI different is the unifying layer on top:
- One result object, one API surface. Every estimator — from
regress()tocallaway_santanna()tocausal_forest()tonotears()— returns aCausalResultwith the same.summary()/.plot()/.to_latex()/.cite()interface. R users juggle 20+ incompatible S3 classes; StatsPAI users juggle one. - Scope no single R or Python package matches. DID + RD + Synth + Matching + DML + Meta-learners + TMLE + Neural Causal + Causal Discovery + Policy Learning + Conformal + Bunching + Spillover + Matrix Completion — all consistent, all under
sp.*. - Agent-native by design. Self-describing schemas (
list_functions(),describe_function(),function_schema()) make StatsPAI the first econometrics toolkit built for LLM-driven research workflows. No other package — in any language — offers this. - Publication pipeline out of the box. Word + Excel + LaTeX + HTML + Markdown export from every estimator, not a separate
modelsummary-style dance.
If a method exists in R, we aim to match or exceed its feature set in Python — and then add what Python can uniquely offer (sklearn integration, JAX/PyTorch backends, agent-native schemas).
| Function | Description | Stata equivalent | R equivalent |
|---|---|---|---|
regress() |
OLS with robust/clustered/HAC SE | reg y x, r / vce(cluster c) |
fixest::feols() |
ivreg() |
IV / 2SLS with first-stage diagnostics | ivregress 2sls |
fixest::feols() with IV |
panel() |
Fixed Effects, Random Effects, Between, FD | xtreg, fe / xtreg, re |
plm::plm() |
heckman() |
Heckman selection model | heckman |
sampleSelection::selection() |
qreg(), sqreg() |
Quantile regression | qreg / sqreg |
quantreg::rq() |
tobit() |
Censored regression (Tobit) | tobit |
censReg::censReg() |
xtabond() |
Arellano-Bond dynamic panel GMM | xtabond |
plm::pgmm() |
glm() |
Generalized Linear Model (6 families × 8 links) | glm |
stats::glm() |
logit(), probit() |
Binary choice with marginal effects | logit / probit |
stats::glm(family=binomial) |
mlogit() |
Multinomial logit | mlogit |
nnet::multinom() |
ologit(), oprobit() |
Ordered logit / probit | ologit / oprobit |
MASS::polr() |
clogit() |
Conditional logit (McFadden) | clogit |
survival::clogit() |
poisson(), nbreg() |
Count data (Poisson, Negative Binomial) | poisson / nbreg |
MASS::glm.nb() |
ppmlhdfe() |
Pseudo-Poisson MLE for gravity models | ppmlhdfe |
fixest::fepois() |
zip_model(), zinb() |
Zero-inflated Poisson / NegBin | zip / zinb |
pscl::zeroinfl() |
hurdle() |
Hurdle (two-part) model | — | pscl::hurdle() |
truncreg() |
Truncated regression (MLE) | truncreg |
truncreg::truncreg() |
fracreg() |
Fractional response (Papke-Wooldridge) | fracreg |
— |
betareg() |
Beta regression | — | betareg::betareg() |
liml() |
LIML (robust to weak IV) | ivregress liml |
AER::ivreg() |
jive() |
Jackknife IV (many instruments) | — | — |
lasso_iv() |
LASSO-selected instruments | — | — |
feols() |
OLS / IV with high-dim fixed effects (pyfixest backend) | reghdfe |
fixest::feols() |
fepois() |
Poisson with high-dim fixed effects | ppmlhdfe |
fixest::fepois() |
feglm() |
GLM with high-dim fixed effects | — | fixest::feglm() |
etable() |
Publication-quality regression tables (LaTeX/Markdown/HTML) | esttab |
fixest::etable() |
sureg() |
Seemingly Unrelated Regression | sureg |
systemfit::systemfit("SUR") |
three_sls() |
Three-Stage Least Squares | reg3 |
systemfit::systemfit("3SLS") |
biprobit() |
Bivariate probit | biprobit |
— |
etregress() |
Endogenous treatment effects | etregress |
— |
gmm() |
General GMM (arbitrary moments) | gmm |
gmm::gmm() |
frontier() |
Stochastic frontier analysis | frontier |
sfa::sfa() |
| Function | Description | Stata equivalent |
|---|---|---|
panel_logit(), panel_probit() |
Panel binary (FE conditional / RE / CRE Mundlak) | xtlogit / xtprobit |
panel_fgls() |
FGLS with heteroskedasticity and AR(1) | xtgls |
interactive_fe() |
Interactive fixed effects (Bai 2009) | — |
panel_unitroot() |
Panel unit root (IPS / LLC / Fisher / Hadri) | xtunitroot |
mixed() |
Linear mixed / multilevel (HLM): unstructured G, 3-level nested, BLUP posterior SEs, Nakagawa–Schielzeth R², caterpillar plot, predict() |
mixed |
melogit(), mepoisson(), meglm() |
Generalised linear mixed models via Laplace approximation (binomial / Poisson / Gaussian) with odds-ratio & IRR tables | melogit / mepoisson / meglm |
icc() |
Intra-class correlation with delta-method 95% CI | estat icc |
lrtest() |
Likelihood-ratio test between nested mixed models with Self–Liang χ̄² boundary correction | lrtest |
| Function | Description | Stata equivalent |
|---|---|---|
cox() |
Cox Proportional Hazards | stcox |
kaplan_meier() |
Kaplan-Meier survival curves | sts graph |
survreg() |
Parametric AFT (Weibull / exponential / log-normal) | streg |
logrank_test() |
Log-rank test for group comparison | sts test |
| Function | Description | Stata equivalent |
|---|---|---|
var() |
Vector Autoregression | var |
granger_causality() |
Granger causality test | vargranger |
irf() |
Impulse response functions | irf graph |
structural_break() |
Bai-Perron structural break test | estat sbsingle |
cusum_test() |
CUSUM parameter stability test | — |
engle_granger() |
Engle-Granger cointegration test | — |
johansen() |
Johansen cointegration (trace / max-eigenvalue) | vecrank |
| Function | Description | Stata equivalent |
|---|---|---|
lpoly() |
Local polynomial regression | lpoly |
kdensity() |
Kernel density estimation | kdensity |
| Function | Description |
|---|---|
randomize() |
Stratified / cluster / block randomization |
balance_check() |
Covariate balance with normalized differences |
attrition_test() |
Differential attrition analysis |
attrition_bounds() |
Lee / Manski bounds under attrition |
optimal_design() |
Optimal sample size / cluster design |
| Function | Description | Stata equivalent |
|---|---|---|
mice() |
Multiple Imputation by Chained Equations | mi impute chained |
mi_estimate() |
Combine estimates via Rubin's rules | mi estimate |
| Function | Description |
|---|---|
mendelian_randomization() |
IVW + MR-Egger + Weighted Median MR |
mr_plot() |
Scatter plot with MR regression lines |
| Function | Description | Reference |
|---|---|---|
blp() |
BLP random-coefficients demand estimation | Berry, Levinsohn & Pakes (1995) |
| Function | Description | Reference |
|---|---|---|
did() |
Auto-dispatching DID (2×2 or staggered) | — |
did_summary() |
One-call robustness comparison across CS/SA/BJS/ETWFE/Stacked | — |
did_summary_plot() |
Forest plot of method-robustness summary | — |
did_summary_to_markdown() / _to_latex() |
Publication-ready tables from did_summary |
— |
did_report() |
One-call bundle: txt + md + tex + png + json into a folder | — |
did_2x2() |
Classic two-group, two-period DID | — |
callaway_santanna() |
Staggered DID with heterogeneous effects | Callaway & Sant'Anna (2021) |
sun_abraham() |
Interaction-weighted event study | Sun & Abraham (2021) |
bacon_decomposition() |
TWFE decomposition diagnostic | Goodman-Bacon (2021) |
honest_did() |
Sensitivity to parallel trends violations | Rambachan & Roth (2023) |
continuous_did() |
Continuous treatment DID (dose-response) | Callaway, Goodman-Bacon & Sant'Anna (2024) |
did_multiplegt() |
DID with treatment switching | de Chaisemartin & D'Haultfoeuille (2020) |
did_imputation() |
Imputation DID estimator | Borusyak, Jaravel & Spiess (2024) |
wooldridge_did() / etwfe() |
Extended TWFE: xvar= (single/multi) + panel= (repeated CS) + cgroup= (never/notyet) |
Wooldridge (2021) |
etwfe_emfx() |
R etwfe::emfx equivalent — simple/group/event/calendar aggregations |
McDermott (2023) |
drdid() |
Doubly robust 2×2 DID (OR + IPW) | Sant'Anna & Zhao (2020) |
stacked_did() |
Stacked event-study DID | Cengiz et al. (2019); Baker, Larcker & Wang (2022) |
ddd() |
Triple-differences (DDD) | Gruber (1994); Olden & Møen (2022) |
cic() |
Changes-in-changes (quantile DID) | Athey & Imbens (2006) |
twfe_decomposition() |
Bacon + de Chaisemartin–D'Haultfoeuille weights | Goodman-Bacon (2021); dCDH (2020) |
distributional_te() |
Distributional treatment effects | Chernozhukov, Fernandez-Val & Melly (2013) |
sp.aggte() |
Unified aggregation for staggered DID (simple/dynamic/group/calendar) with Mammen multiplier-bootstrap uniform bands | Callaway & Sant'Anna (2021) §4; Mammen (1993) |
sp.cs_report() |
One-call Callaway–Sant'Anna report: estimation + four aggregations + pre-trend test + Rambachan–Roth breakdown M* | CS2021 + RR2023 |
sp.ggdid() |
aggte() visualiser with uniform-band overlay |
mirrors R did::ggdid |
CSReport.plot() |
2×2 summary figure (event study / θ(g) / θ(t) / RR breakdown) | — |
CSReport.to_markdown() |
GitHub-Flavoured Markdown export of the full report | — |
CSReport.to_latex() |
Booktabs LaTeX fragment, jinja2-free | — |
CSReport.to_excel() |
Six-sheet Excel workbook | — |
All algorithms below are reimplemented from the original papers — no wrappers, no runtime dependencies on upstream DID packages.
| Feature | StatsPAI | csdid (Py) |
differences (Py) |
R did |
|---|---|---|---|---|
| Callaway–Sant'Anna ATT(g,t) with DR / IPW / REG | ✅ | ✅ | ✅ | ✅ |
| Never-treated / not-yet-treated control group | ✅ | ✅ | ✅ | ✅ |
Anticipation (anticipation=δ) |
✅ | ✅ | — | ✅ |
Repeated cross-sections (panel=False) |
✅ | ✅ | partial | ✅ |
aggte: simple / dynamic / group / calendar |
✅ | ✅ | ✅ | ✅ |
| Mammen multiplier bootstrap, uniform sup-t bands | ✅ | ✅ | — | ✅ |
balance_e / min_e / max_e |
✅ | ✅ | partial | ✅ |
| Sun–Abraham IW with Liang–Zeger cluster SE | ✅ | — | ✅ | via fixest::sunab |
| Borusyak–Jaravel–Spiess imputation + pre-trend Wald | ✅ | — | — | via didimputation |
| de Chaisemartin–D'Haultfoeuille switch-on-off | ✅ | — | — | via DIDmultiplegtDYN |
| dCDH joint placebo Wald + avg. cumulative effect | ✅ | — | — | ✅ (v2) |
| Rambachan–Roth sensitivity + breakdown M* | ✅ | — | — | via HonestDiD |
cs ⇄ aggte ⇄ honest_did pipeline (single object) |
✅ | partial | partial | partial |
One-call report card (cs_report) |
✅ | — | — | via summary() |
| Markdown / LaTeX / Excel report export | ✅ | — | — | partial |
save_to= one-call bundle (txt + md + tex + xlsx + png) |
✅ | — | — | — |
CSReport.plot() 2×2 summary figure |
✅ | — | — | — |
| Function | Description | Reference |
|---|---|---|
rdrobust() |
Sharp/Fuzzy RD with robust bias-corrected inference | Calonico, Cattaneo & Titiunik (2014) |
rdplot() |
RD visualization with binned scatter | — |
rddensity() |
McCrary density manipulation test | McCrary (2008) |
rdmc() |
Multi-cutoff RD | Cattaneo et al. (2024) |
rdms() |
Geographic / multi-score RD | Keele & Titiunik (2015) |
rkd() |
Regression Kink Design | Card et al. (2015) |
| Function | Description | Stata equivalent |
|---|---|---|
match() |
PSM, Mahalanobis, CEM with balance diagnostics | psmatch2 / cem |
ebalance() |
Entropy balancing | ebalance |
| Function | Description | Reference |
|---|---|---|
synth() |
Abadie-Diamond-Hainmueller SCM | Abadie et al. (2010) |
sdid() |
Synthetic Difference-in-Differences | Arkhangelsky et al. (2021) |
| Placebo inference, gap plots, weight tables, RMSE plots | — | — |
| Function | Description | Reference |
|---|---|---|
dml() |
Double/Debiased ML (PLR + IRM) with cross-fitting | Chernozhukov et al. (2018) |
causal_forest() |
Causal Forest for heterogeneous treatment effects | Wager & Athey (2018) |
deepiv() |
Deep IV neural network approach | Hartford et al. (2017) |
metalearner() |
S/T/X/R/DR-Learner for CATE estimation | Kunzel et al. (2019), Kennedy (2023) |
tmle() |
Targeted Maximum Likelihood Estimation | van der Laan & Rose (2011) |
aipw() |
Augmented Inverse-Probability Weighting | — |
| Function | Description | Reference |
|---|---|---|
tarnet() |
Treatment-Agnostic Representation Network | Shalit et al. (2017) |
cfrnet() |
Counterfactual Regression Network | Shalit et al. (2017) |
dragonnet() |
Dragon Neural Network for CATE | Shi et al. (2019) |
| Function | Description | Reference |
|---|---|---|
notears() |
DAG learning via continuous optimization | Zheng et al. (2018) |
pc_algorithm() |
Constraint-based causal graph learning | Spirtes et al. (2000) |
| Function | Description | Reference |
|---|---|---|
policy_tree() |
Optimal treatment assignment rules | Athey & Wager (2021) |
policy_value() |
Policy value evaluation | — |
| Function | Description | Reference |
|---|---|---|
conformal_cate() |
Distribution-free prediction intervals for ITE | Lei & Candes (2021) |
bcf() |
Bayesian Causal Forest (separate mu/tau) | Hahn, Murray & Carvalho (2020) |
| Function | Description | Reference |
|---|---|---|
dose_response() |
Continuous treatment dose-response curve (GPS) | Hirano & Imbens (2004) |
multi_treatment() |
Multi-valued treatment AIPW | Cattaneo (2010) |
| Function | Description | Reference |
|---|---|---|
lee_bounds() |
Sharp bounds under sample selection | Lee (2009) |
manski_bounds() |
Worst-case bounds (no assumption / MTR / MTS) | Manski (1990) |
| Function | Description | Reference |
|---|---|---|
spillover() |
Direct + spillover + total effect decomposition | Hudgens & Halloran (2008) |
| Function | Description | Reference |
|---|---|---|
g_estimation() |
Multi-stage optimal DTR via G-estimation | Robins (2004) |
| Function | Description | Reference |
|---|---|---|
bunching() |
Kink/notch bunching estimator with elasticity | Kleven & Waseem (2013) |
| Function | Description | Reference |
|---|---|---|
mc_panel() |
Causal panel data via nuclear-norm matrix completion | Athey et al. (2021) |
| Function | Description | Stata/R equivalent |
|---|---|---|
causal_impact() |
Bayesian structural time-series | R CausalImpact |
mediate() |
Mediation analysis (ACME/ADE) | medeff / R mediation |
bartik() |
Shift-share IV with Rotemberg weights | bartik_weight |
| Function | Description | Stata equivalent |
|---|---|---|
margins() |
Average marginal effects (AME/MEM) | margins, dydx(*) |
marginsplot() |
Marginal effects visualization | marginsplot |
test() |
Wald test for linear restrictions | test x1 = x2 |
lincom() |
Linear combinations with inference | lincom x1 + x2 |
| Function | Description | Reference |
|---|---|---|
oster_bounds() |
Coefficient stability bounds | Oster (2019) |
sensemakr() |
Sensitivity to omitted variables | Cinelli & Hazlett (2020) |
mccrary_test() |
Density discontinuity test | McCrary (2008) |
hausman_test() |
FE vs RE specification test | Hausman (1978) |
anderson_rubin_test() |
Weak instrument robust inference + AR confidence set | Anderson & Rubin (1949) |
effective_f_test() |
Heteroskedasticity-robust effective F (HC1) | Olea & Pflueger (2013) |
tF_critical_value() |
Adjusted t-ratio critical value (valid under weak IV) | Lee, McCrary, Moreira & Porter (2022, AER) |
evalue() |
E-value sensitivity to unmeasured confounding | VanderWeele & Ding (2017) |
het_test() |
Breusch-Pagan / White heteroskedasticity | — |
reset_test() |
Ramsey RESET specification test | — |
vif() |
Variance Inflation Factor | — |
diagnose() |
General model diagnostics | — |
| Function | Description |
|---|---|
recommend() |
Given data + research question → recommends estimators with reasoning, generates workflow, provides .run() |
compare_estimators() |
Runs multiple methods (OLS, matching, IPW, DML, ...) on same data, reports agreement diagnostics |
assumption_audit() |
One-call test of ALL assumptions for any method, with pass/fail/remedy for each |
sensitivity_dashboard() |
Multi-dimensional sensitivity analysis (sample, outliers, unobservables) with stability grade |
pub_ready() |
Journal-specific publication readiness checklist (Top 5 Econ, AEJ, RCT) |
replicate() |
Built-in famous datasets (Card 1995, LaLonde 1986, Lee 2008) with replication guides |
| Function | Description | R/Stata equivalent |
|---|---|---|
spec_curve() |
Specification Curve / Multiverse Analysis | R specr (limited) / Stata: none |
robustness_report() |
Automated robustness battery (SE variants, winsorize, trim, add/drop controls, subsamples) | None |
subgroup_analysis() |
Heterogeneity analysis with forest plot + interaction Wald test | None (manual in both) |
| Function | Description |
|---|---|
wild_cluster_bootstrap() |
Wild cluster bootstrap (Cameron, Gelbach & Miller 2008) |
ri_test() |
Randomization inference / Fisher exact test |
| Function | Description |
|---|---|
cate_summary(), cate_by_group() |
CATE distribution summaries |
cate_plot(), cate_group_plot() |
CATE visualization |
gate_test() |
Group Average Treatment Effect test |
blp_test() |
Best Linear Projection test |
compare_metalearners() |
Compare S/T/X/R/DR-Learner estimates |
| Function | Description | Formats |
|---|---|---|
modelsummary() |
Multi-model comparison tables | Text, LaTeX, HTML, Word, Excel, DataFrame |
outreg2() |
Stata-style regression table export | Excel, LaTeX, Word |
sumstats() |
Summary statistics (Table 1) | Text, LaTeX, HTML, Word, Excel, DataFrame |
balance_table() |
Pre-treatment balance check | Text, LaTeX, HTML, Word, Excel, DataFrame |
tab() |
Cross-tabulation with chi-squared / Fisher | Text, LaTeX, Word, Excel, DataFrame |
coefplot() |
Coefficient forest plot across models | matplotlib Figure |
binscatter() |
Binned scatter with residualization | matplotlib Figure |
set_theme() |
Publication themes ('academic', 'aea', 'minimal', 'cn_journal') |
— |
interactive() |
WYSIWYG plot editor with 29 themes & auto code generation | Jupyter ipywidgets |
Every result object has:
result.summary() # Formatted text summary
result.plot() # Appropriate visualization
result.to_latex() # LaTeX table
result.to_docx() # Word document
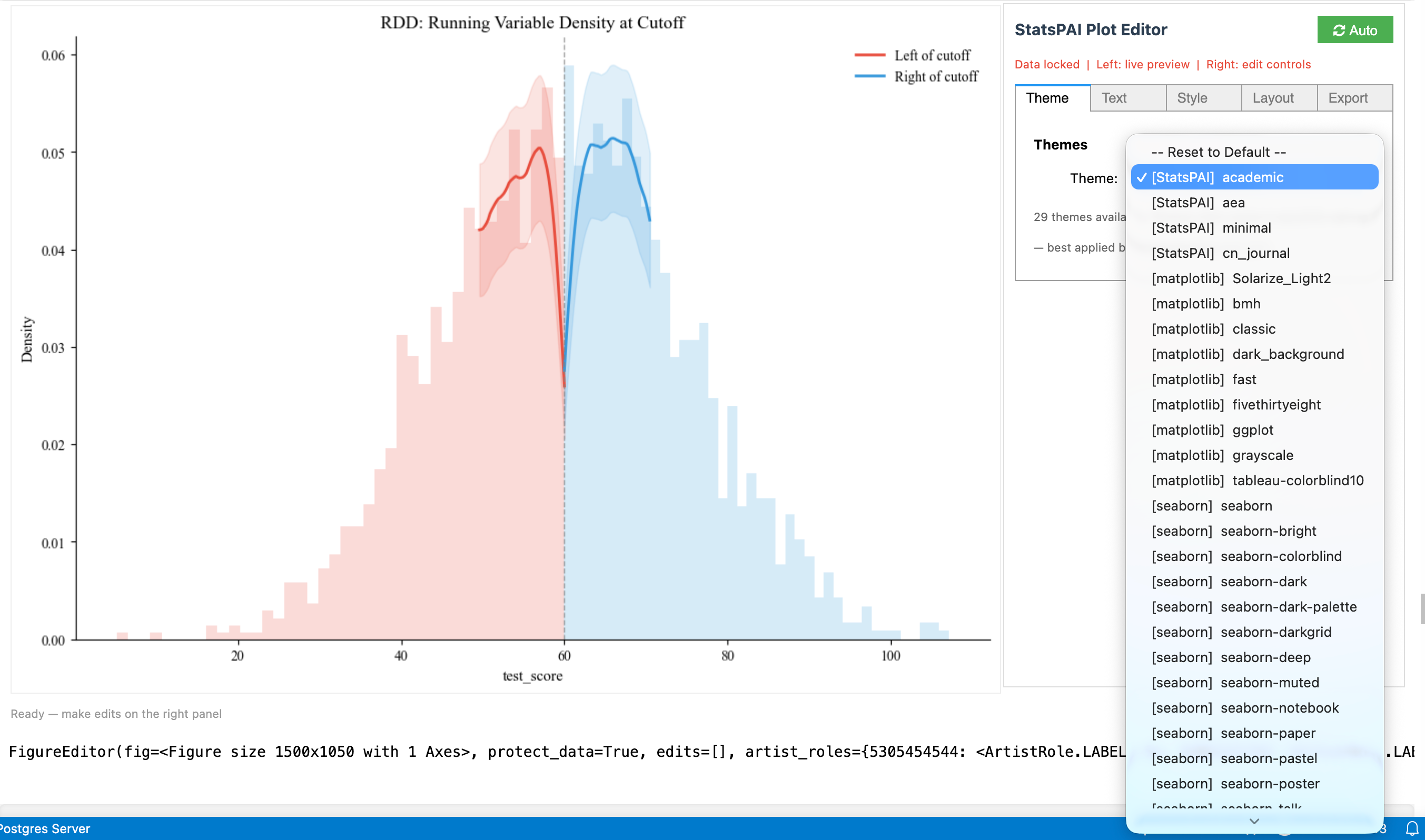
result.cite() # BibTeX citation for the methodStata users know the Graph Editor: double-click a figure to enter a WYSIWYG editing interface — drag fonts, change colors, adjust layout. This has been a Stata-exclusive experience. In Python, matplotlib produces static images — changing a title font size means editing code and re-running.
sp.interactive(fig) turns any matplotlib figure into a live editing panel — figure preview on the left, property controls on the right, just like Stata's Graph Editor. But it does two things Stata can't:
-
29 academic themes, one-click switching. From AER journal style to ggplot, FiveThirtyEight, dark presentation mode — select and see the result instantly. Stata's
schemerequires regenerating the plot; here it's real-time. -
Every edit auto-generates reproducible Python code. Adjust title size, change colors, add annotations in the GUI — the editor records each operation as standard matplotlib code (
ax.set_title(...),ax.spines[...].set_visible(...)). Copy with one click, paste into your script, and it reproduces exactly. Stata's Graph Editor cannot export edits to do-file commands.
Five tabs cover all editing needs: Theme (29 themes) · Text (titles, labels, fonts) · Style (line colors, widths, markers) · Layout (spines, grid, figure size, legend, axis limits) · Export (save, undo/redo, reset).
Auto/Manual rendering modes: Auto refreshes the preview on every change; Manual batches edits for a single Apply — useful for large figures or slow machines.
import statspai as sp
result = sp.did(df, y='wage', treat='policy', time='year')
fig, ax = result.plot()
editor = sp.interactive(fig) # opens the editor
# After editing in the GUI:
editor.copy_code() # prints reproducible Python code| Function | Description | Stata equivalent |
|---|---|---|
label_var(), label_vars() |
Variable labeling | label var |
describe() |
Data description | describe |
pwcorr() |
Pairwise correlation with significance stars | pwcorr, star(.05) |
winsor() |
Winsorization | winsor2 |
read_data() |
Multi-format data reader | use / import |
pip install statspaiWith optional dependencies:
pip install statspai[plotting] # matplotlib, seaborn
pip install statspai[fixest] # pyfixest for high-dimensional FERequirements: Python >= 3.9
Core dependencies: NumPy, SciPy, Pandas, statsmodels, scikit-learn, linearmodels, patsy, openpyxl, python-docx
import statspai as sp
# --- Estimation ---
r1 = sp.regress("wage ~ education + experience", data=df, robust='hc1')
r2 = sp.ivreg("wage ~ (education ~ parent_edu) + experience", data=df)
r3 = sp.did(df, y='wage', treat='policy', time='year', id='worker')
r4 = sp.rdrobust(df, y='score', x='running_var', c=0)
r5 = sp.dml(df, y='wage', treat='training', covariates=['age', 'edu', 'exp'])
r6 = sp.causal_forest("y ~ treatment | x1 + x2 + x3", data=df)
# --- Post-estimation ---
sp.margins(r1, data=df) # Marginal effects
sp.test(r1, "education = experience") # Wald test
sp.oster_bounds(df, y='wage', treat='education', controls=['experience'])
# --- Tables (to Word / Excel / LaTeX) ---
sp.modelsummary(r1, r2, output='table2.docx')
sp.outreg2(r1, r2, r3, filename='results.xlsx')
sp.sumstats(df, vars=['wage', 'education', 'age'], output='table1.docx')
# --- Robustness (unique to StatsPAI) ---
sp.spec_curve(df, y='wage', x='education',
controls=[[], ['experience'], ['experience', 'female']],
se_types=['nonrobust', 'hc1']).plot()
sp.robustness_report(df, formula="wage ~ education + experience",
x='education', extra_controls=['female'],
winsor_levels=[0.01, 0.05]).plot()
sp.subgroup_analysis(df, formula="wage ~ education + experience",
x='education',
by={'Gender': 'female', 'Region': 'region'}).plot()| Advantage | Detail |
|---|---|
| Unified API | One package, one import, consistent .summary() / .plot() / .to_latex() across all methods. Stata requires paid add-ons; R requires 20+ packages with different interfaces. |
| Modern ML causal methods | DML, Causal Forest, Meta-Learners (S/T/X/R/DR), TMLE, DeepIV, TARNet/CFRNet/DragonNet, Policy Trees — all in one place. Stata has almost none of these. R has them scattered across incompatible packages. |
| Robustness automation | spec_curve(), robustness_report(), subgroup_analysis() — no manual re-running. Neither Stata nor R offers this out-of-the-box. |
| Free & open source | MIT license, $0. Stata costs $695–$1,595/year. |
| Python ecosystem | Integrates naturally with pandas, scikit-learn, PyTorch, Jupyter, cloud pipelines. |
| Auto-citations | Every causal method has .cite() returning the correct BibTeX. Neither Stata nor R does this. |
| Interactive Plot Editor | sp.interactive() — Stata Graph Editor-style GUI in Jupyter with 29 themes and auto-generated reproducible code. Stata's Graph Editor can't export edits to do-file; R has no equivalent. |
| Advantage | Detail |
|---|---|
| Battle-tested at scale | 40+ years of production use in economics. Edge cases are well-handled. |
| Speed on very large datasets | Stata's compiled C backend is faster for simple OLS/FE on datasets with millions of rows. |
| Survey data & complex designs | svy: prefix, stratification, clustering — Stata's survey support is unmatched. |
| Mature documentation | Every command has a PDF manual with worked examples. Community is massive. |
| Journal acceptance | Referees in some fields trust Stata output by default. |
| Advantage | Detail |
|---|---|
| Cutting-edge methods | New econometric methods (e.g., fixest, did2s, HonestDiD) often appear in R first. |
ggplot2 visualization |
R's grammar of graphics is more flexible than matplotlib for complex figures. |
modelsummary |
R's modelsummary is the gold standard for regression tables — StatsPAI's is close but not yet identical. |
| CRAN quality control | R packages go through peer review. Python packages vary in quality. |
| Spatial econometrics | spdep, spatialreg |
450+ public functions/classes
Regression: regress, ivreg, glm, logit, probit, mlogit, ologit, poisson, nbreg, ppmlhdfe,
tobit, heckman, qreg, truncreg, fracreg, betareg, sureg, three_sls, gmm
IV Advanced: liml, jive, lasso_iv
Panel: panel, panel_logit, panel_probit, panel_fgls, interactive_fe, xtabond, mixed
DID: did, callaway_santanna, sun_abraham, bacon_decomposition, honest_did,
continuous_did, did_multiplegt, did_imputation, stacked_did
RD: rdrobust, rdplot, rddensity, rdmc, rdms, rkd
Matching: match, ebalance, ipw, aipw
Synth: synth, sdid, gsynth, augsynth, staggered_synth, conformal_synth
ML Causal: dml, causal_forest, deepiv, metalearner, tmle
Neural: tarnet, cfrnet, dragonnet
Spatial: sar, sem, sdm, slx, sac, sar_gmm, sem_gmm, sarar_gmm, ← NEW v0.8
moran, geary, getis_ord_g, join_counts, lm_tests, impacts,
gwr, mgwr, gwr_bandwidth, spatial_panel,
queen_weights, rook_weights, knn_weights, distance_band, kernel_weights
Discovery: notears, pc_algorithm, lingam, ges ← NEW v0.8
Policy: policy_tree, policy_value
Survival: cox, kaplan_meier, survreg, logrank_test, cox_frailty, aft ← NEW v0.8
Time Series: var, granger_causality, irf, structural_break, johansen,
local_projections, garch, arima, bvar ← NEW v0.8
Nonparametric: lpoly, kdensity
Experimental: randomize, balance_check, attrition_test, optimal_design
Matching: match, ebalance, optimal_match, cardinality_match ← NEW v0.8
Decomposition: oaxaca, gelbach, rifreg, rif_decomposition ← NEW v0.8
Imputation: mice, mi_estimate
Frontier: frontier (stochastic frontier analysis)
Structural: blp (BLP demand estimation)
Survey: svydesign, svymean, svytotal, svyglm, rake, linear_calibration ← NEW v0.8
MR: mendelian_randomization, mr_ivw, mr_egger, mr_median
Mediation: mediate, mediate_sensitivity ← NEW v0.8
RD: rdrobust, rdplot, rddensity, rdmc, rdms, rdpower, rdsampsi ← NEW v0.8
Smart Workflow: recommend, compare_estimators, assumption_audit,
sensitivity_dashboard, pub_ready, replicate
Output: modelsummary, outreg2, sumstats, balance_table, tab, coefplot, binscatter
Plot Editor: interactive (WYSIWYG editor), set_theme (29 academic themes)
24 commits since 0.9.2 (includes the GLMM hardening pillar originally tagged as 0.9.4, folded into 0.9.3 before tag). See CHANGELOG for full detail.
sp.frontier. A latent Jondrow-posterior sign error in all prior versions (≤ 0.9.2) produced systematically biased efficiency scores; the dist='exponential' path additionally returned NaN for unit efficiency. Re-run any prior frontier analyses.
1. Stochastic Frontier — full Stata/R parity and beyond. sp.frontier + sp.xtfrontier rewritten (~2,700 LOC).
- Cross-sectional: heteroskedastic inefficiency
usigma=[...](Caudill-Ford-Gropper 1995), heteroskedastic noisevsigma=[...](Wang 2002), inefficiency determinantsemean=[...](Kumbhakar-Ghosh-McGuckin 1991); Battese-Coelli (1988)E[exp(-u)|ε]alongside JLMS; LR mixed-χ̄² test (Kodde-Palm 1986); parametric-bootstrap unit-efficiency CIs. - Panel: Pitt-Lee (1981) time-invariant; Battese-Coelli (1992) time-decay; Battese-Coelli (1995) full-flexibility; Greene (2005) TFE/TRE; Dhaene-Jochmans (2015) split-panel jackknife bias correction via
xtfrontier(..., model='tfe', bias_correct=True). - New advanced frontiers:
sp.zisf— Zero-Inefficiency SFA mixture (Kumbhakar-Parmeter-Tsionas 2013);sp.lcsf— 2-class Latent-Class SFA (Orea-Kumbhakar 2004 / Greene 2005). - Productivity:
sp.malmquist— Färe-Grosskopf-Lindgren-Roos (1994) Malmquist TFP index with M = EC × TC decomposition;sp.translog_design— Cobb-Douglas → Translog design-matrix helper. - Inference + post-estimation:
vce='opg' / 'robust' / 'cluster' / 'bootstrap'with Monte-Carlo coverage check; conditionalpredict(), returns-to-scale,usigma/emeanmarginal effects, metafrontier.
2. Multilevel / Mixed-Effects — sp.multilevel rewritten. From a 400-LOC two-level single file to a ~2,700-LOC sub-package with lme4/Stata mixed parity. sp.mixed now defaults to unstructured random-effect covariance, supports three-level nested models (group=['school','class']), BLUP posterior SEs, Nakagawa-Schielzeth R², caterpillar plots. New top-level: sp.melogit / sp.mepoisson / sp.meglm (Laplace GLMMs), sp.icc (delta-method CI), sp.lrtest (Self-Liang χ̄² boundary correction). Verified against statsmodels.MixedLM to 4 decimal places.
3. GLMM hardening — AGHQ + Gamma / NegBin / Ordinal. Closes the three GLMM gaps flagged in the multilevel self-audit. Adaptive Gauss-Hermite quadrature via new nAGQ argument: nAGQ=1 reduces exactly to Laplace (verified 1e-10); nAGQ>1 matches Stata intpoints(7) / R lme4::glmer(nAGQ=7) accuracy on small clusters with binary or other non-Gaussian outcomes. Wired into all five families (Gaussian / Binomial / Poisson / Gamma / NegBin) plus meologit. New families: sp.megamma (Gamma GLMM, log link, ML-estimated dispersion log φ, Fisher-scoring IRLS), sp.menbreg (NB-2 Var = μ + αμ², reduces to Poisson as α → 0), sp.meologit (random-effects ordinal logit with K−1 reparameterised thresholds κ_1, log(κ_2−κ_1), … for unconditional ordering). Cross-family AIC comparability: Poisson and Binomial log-likelihoods now include full normalisation constants (-log(y!) / log-binomial-coefficient) so mepoisson vs menbreg AIC comparisons are unbiased.
4. Econometric Trinity — three new P0 pillars (~1,170 LOC).
sp.dml(model='pliv', instrument=...)— Partially Linear IV (Chernozhukov et al. 2018, §4.2) with Neyman-orthogonal score and cross-fittedg/m/rnuisances; influence-function SEs.sp.mixlogit— Random-coefficient MNL via simulated ML with Halton draws. Normal / log-normal / triangular mixing; diagonal or Cholesky covariance; panel repeated-choice; OPG-sandwich SEs. Python's first feature-complete implementation.sp.ivqreg— Chernozhukov-Hansen IV quantile regression via inverse-QR profile. Grid + Brent (scalar endogenous); BFGS onb̂(α)(multi-dim); pairs-bootstrap SEs.
5. Smart workflow — posterior verification. sp.verify / sp.verify_benchmark — aggregates bootstrap stability + placebo pass rate + subsample agreement into a verify_score ∈ [0, 100] for any sp.recommend() output. Opt-in via recommend(verify=True); zero overhead when off.
Quality bar. Multilevel passed oracle + code-reviewer audit (4 BLOCKER + 5 HIGH fixed); econ-trinity passed self-audit (4 BLOCKER + 7 HIGH fixed); frontier self-audit fixed Mills-tail, TVD-loop, cost-panel, summary-dump issues. GLMM hardening added 18 new tests (TestAGHQ × 7, TestMEGamma × 3, TestMENegBin × 3, TestMEOLogit × 5) on top of the 35 prior multilevel tests. Test count: 93/93 frontier, 53/53 multilevel (incl. GLMM), 18/18 smart, 10/10 econ-trinity pass.
Meta. Author attribution corrected from "Bryce Wang" to "Biaoyue Wang" in pyproject.toml, __author__, English/Chinese READMEs, docs/index.md, and mkdocs.yml (JOSS paper.md was already correct).
Release focus: statspai.decomposition. 18 first-class decomposition methods across 13 modules (~6,200 LOC, 54 tests) — Python's first (and most complete) implementation of the full decomposition toolkit spanning mean, distributional, inequality, demographic, and causal decomposition. Occupies the previously empty Python high-ground where only one unmaintained 2018-vintage PyPI package existed.
18 methods (30 aliases) under sp.decompose():
- Mean —
oaxaca(Blinder-Oaxaca threefold with 5 reference coefficients: Neumark 1988, Cotton 1988, Reimers 1983, group A, group B),gelbach(Gelbach 2016 sequential OVB),fairlie(nonlinear logit/probit),bauer_sinning/yun_nonlinear(detailed nonlinear) - Distributional —
rifreg/rif_decomposition(FFL 2009),ffl_decompose(FFL 2018 two-step),dfl_decompose(DiNardo-Fortin-Lemieux 1996 reweighting),machado_mata(MM 2005),melly_decompose(Melly 2005),cfm_decompose(Chernozhukov-Fernández-Val-Melly 2013) - Inequality —
subgroup_decompose(Theil T/L, GE(α), Dagum Gini, Atkinson, CV²),shapley_inequality(Shorrocks 2013),source_decompose(Lerman-Yitzhaki 1985) - Demographic —
kitagawa_decompose(1955),das_gupta(1993 multi-factor symmetric) - Causal —
gap_closing(Lundberg 2021 regression/IPW/AIPW),mediation_decompose(VanderWeele 2014 NDE/NIE),disparity_decompose(Jackson-VanderWeele 2018)
Quality bar:
- Closed-form influence functions for Theil T / Theil L / Atkinson (no O(n²) numerical fallback)
- Weighted O(n log n) Dagum Gini via sorted-ECDF pairwise-MAD identity
- Cross-method consistency tests (
test_dfl_ffl_mean_agree,test_mm_melly_cfm_aligned_reference,test_dfl_mm_reference_convention_opposite) - Numerical identity checks: FFL four-part sum, weighted Gini RIF
E_w[RIF] = G - Logit non-convergence surfaces as
RuntimeWarning; bootstrap failure rate >5% warns - 54 decomposition tests + core refactor consolidation:
decomposition/_common.pyhostsinfluence_function(y, stat, tau, w)— the canonical 9-statistic RIF kernel (rif.rif_valuespublic API expands from 3 to 9 statistics)
Unified entry point:
import statspai as sp
result = sp.decompose(method='ffl', data=df, y='log_wage',
group='female', x=['education', 'experience'],
stat='quantile', tau=0.5)
result.summary(); result.plot(); result.to_latex()Release focus: statspai.rd. 18+ RD estimators, diagnostics, and inference methods across 14 modules (~10,300 LOC) — the most feature-complete RD package in Python, R, or Stata. Full machinery behind CCT, Cattaneo-Jansson-Ma density tests, Armstrong-Kolesar honest CIs, Cattaneo-Titiunik-Vazquez-Bare local randomization, Cattaneo-Titiunik-Yu boundary (2D) RD, and Angrist-Rokkanen external validity — all under sp.*.
Core estimation:
rdrobust— sharp / fuzzy / kink RD with bias-corrected robust inference (CCT 2014); covariate-adjusted local polynomial (CCFT 2019)rd2d— boundary discontinuity / 2D RD (Cattaneo, Titiunik & Yu 2025)rkd— Regression Kink Design (Card, Lee, Pei & Weber 2015)rdit— Regression Discontinuity in Time (Hausman & Rapson 2018)rdmc,rdms— multi-cutoff and multi-score RD
Bandwidth selection: rdbwselect with mserd, msetwo, cerrd, cercomb1, cercomb2 (Imbens-Kalyanaraman 2012; CCF 2020).
Inference:
rd_honest— honest CIs with worst-case bias bound (Armstrong-Kolesar 2018, 2020)rdrandinf,rdwinselect,rdsensitivity— local randomization inference (Cattaneo-Frandsen-Titiunik 2015)rdrbounds— Rosenbaum sensitivity bounds for hidden selection
Heterogeneous treatment effects: rdhte (CATE via fully interacted local linear), rdbwhte, rd_forest, rd_boost, rd_lasso.
External validity & extrapolation: rd_extrapolate (Angrist-Rokkanen 2015), rd_multi_extrapolate (CKTV 2024).
Diagnostics & one-click dashboard: rdsummary (rdrobust + density test + bandwidth sensitivity + placebo cutoffs + covariate balance), rdplot (IMSE-optimal binning), rddensity (CJM 2020), rdbalance, rdplacebo.
Power analysis: rdpower, rdsampsi.
Refactor — rd/_core.py: 5-sprint refactor consolidated shared low-level primitives duplicated across 9 RD files into one 191-line canonical private module (_kernel_fn, _kernel_constants, _local_poly_wls, _sandwich_variance). 253 lines of duplicated math → 191 lines of canonical implementation. 97 RD tests pass with zero regression.
Also in 0.9.1:
synth/_core.py— simplex weight solver consolidated from 6 duplicate implementations; analytic Jacobian now available to all six callers (~3-5× speedup)- Bug fixes: density test CJM (2020) implementation + DGP helpers + validation tests; 3 critical + 3 high-priority bugs from comprehensive RD code review;
_ols_fitsingular matrix fallback
Release focus: statspai.synth. 20 SCM methods + 6 inference strategies + full research workflow (compare / power / sensitivity / one-click reports), all behind the unified sp.synth(method=...) dispatcher. No competing package in Python, R, or Stata offers this breadth.
Seven new SCM estimators:
| Method | Reference |
|---|---|
bayesian_synth |
Dirichlet-prior MCMC with full posterior credible intervals (Vives & Martinez 2024) |
bsts_synth / causal_impact |
Bayesian Structural Time Series via Kalman filter/smoother (Brodersen et al. 2015) |
penalized_synth (penscm) |
Pairwise discrepancy penalty (Abadie & L'Hour 2021, JASA) |
fdid |
Forward DID with optimal donor subset selection (Li 2024) |
cluster_synth |
K-means / spectral / hierarchical donor clustering (Rho 2024) |
sparse_synth |
L1 / constrained-LASSO / joint V+W (Amjad, Shah & Shen 2018, JMLR) |
kernel_synth + kernel_ridge_synth |
RKHS / MMD-based nonlinear matching |
Previous methods — classic, penalized, demeaned, unconstrained, augmented (ASCM), SDID, gsynth, staggered, MC, discos, multi-outcome, scpi — remain with bug fixes.
Research workflow:
synth_compare(df, ...)— run every method at once, tabular + graphical comparisonsynth_recommend(df, ...)— auto-select best estimator by pre-fit + robustnesssynth_report(result, format='markdown'|'latex'|'text')— one-click publication-ready reportsynth_power(df, effect_sizes=[...])— first power-analysis tool for SCM designssynth_mde(df, target_power=0.8)— minimum detectable effectsynth_sensitivity(result)— LOO + time placebos + donor sensitivity + RMSPE filtering- Canonical datasets:
california_tobacco(),german_reunification(),basque_terrorism()
Release-blocker fixes (5-parallel-agent code review — correctness / numerics / API / perf / docs):
- ASCM correction formula —
augsynthnow follows Ben-Michael, Feller & Rothstein (2021) Eq. 3 per-period ridge bias(Y1_pre − Y0'γ) @ β(T0, T1), replacing the scalar mean-residual placeholder;_ridge_fitRHS bug also fixed - Bayesian likelihood scale — covariate rows z-scored to pooled pre-outcome SD before concatenation
- Bayesian MCMC Jacobian — missing
log(σ′/σ)correction for log-normal random-walk proposal on σ added to MH acceptance ratio - BSTS Kalman filter — innovation variance floored at
1e-12; RTS smootherinv → solve + pinvfallback on near-singular predicted covariance - gsynth factor estimation — four
np.linalg.inv→np.linalg.lstsq(robust to rank-deficientF'F/L'L) - Dispatcher
**kwargsleakage fixed foraugsynthandkernel_ridgeplacebo forwarding - Cross-method API consistency —
sdid()accepts canonicaloutcome / treated_unit / treatment_time(legacy aliases retained)
Tests & validation: 144 synth tests passing (new: 12-method cross-method consistency benchmark). Full suite: 1481 passed, 4 skipped, 0 failed. New guide: docs/guides/synth.md with a 20-method decision table.
60+ new functions, 450+ total API, 1,230+ tests passing. Largest release in StatsPAI history.
Spatial Econometrics (NEW — 38 API symbols, 3,178 LOC, 69 tests):
- Weights:
W(sparse CSR),queen_weights,rook_weights,knn_weights,distance_band,kernel_weights,block_weights - ESDA:
moran(global + local LISA),geary,getis_ord_g,getis_ord_local,join_counts,moran_plot,lisa_cluster_map - ML Regression:
sar,sem,sdm,slx,sac(SARAR) — sparse-aware, scales to N=100K - GMM Regression:
sar_gmm,sem_gmm,sarar_gmm(Kelejian-Prucha 1998/1999, het-robust) - Diagnostics:
lm_tests(Anselin 1988 — LM-err/LM-lag/Robust-LM/SARMA),moran_residuals - Effects:
impacts(LeSage-Pace 2009 direct/indirect/total with simulated SE) - GWR:
gwr,mgwr(Multiscale GWR),gwr_bandwidth(AICc/CV golden-section) - Spatial Panel:
spatial_panel(SAR-FE / SEM-FE / SDM-FE, entity + twoways) - Cross-validated: Columbus SAR/SEM rtol<1e-7 vs PySAL spreg; Georgia GWR bit-identical vs mgwr
Time Series (4 new estimators):
local_projections— Jordà (2005) horizon-by-horizon IRF with Newey-West HACgarch— GARCH(p,q) volatility model, MLE, multi-step forecastarima— ARIMA/SARIMAX with auto (p,d,q) selection via AICc gridbvar— Bayesian VAR with Minnesota (Litterman) prior, closed-form posterior
Causal Discovery (2 new algorithms):
lingam— DirectLiNGAM (Shimizu 2011), bit-identical match vs lingam packageges— Greedy Equivalence Search (Chickering 2002), BIC-based CPDAG learning
Matching (2 new methods):
optimal_match— global 1:1 matching via Hungarian algorithm (min total Mahalanobis distance)cardinality_match— Zubizarreta (2014) LP-based matching with SMD balance constraints
Decomposition & Mediation:
rifreg— RIF regression (Firpo-Fortin-Lemieux 2009), unconditional quantile partial effectsrif_decomposition— RIF Oaxaca-Blinder for distributional statisticsmediate_sensitivity— Imai-Keele-Yamamoto (2010) ρ-sensitivity analysis
RD & Design:
rdpower,rdsampsi— power calculations for RD designs (Cattaneo et al. 2019)
Survey:
rake— iterative proportional fitting (Deming-Stephan) for survey calibrationlinear_calibration— Deville-Särndal (1992) chi-squared distance calibration
Survival (2 new models):
cox_frailty— Cox proportional hazards with shared gamma frailtyaft— Accelerated Failure Time (exponential/Weibull/lognormal/loglogistic)
ML-Causal (GRF extensions):
CausalForest.variable_importance()— permutation-based feature importance for CATECausalForest.best_linear_projection()— BLP heterogeneity test (Chernozhukov et al. 2020)CausalForest.ate(),.att()— convenience accessors- Bugfix: honest leaf values now correctly vary per-leaf (was overwriting all leaves)
Infrastructure:
- OLS/IV
predict(data, what='confidence'|'prediction')— out-of-sample with intervals - Pre-release code review: 3 critical + 2 high-priority bugs fixed before release
30 new modules, 390+ public API, 860+ tests passing, 83K+ lines of code.
New Regression & GLM:
glm()(6 families × 8 links),logit(),probit(),cloglog(),mlogit(),ologit(),oprobit(),clogit()poisson(),nbreg(),ppmlhdfe()(gravity model),zip_model(),zinb(),hurdle()truncreg(),fracreg(),betareg(),biprobit(),etregress()liml(),jive(),lasso_iv()(advanced IV),sureg(),three_sls(),gmm()(general GMM)
New Panel & Multilevel:
panel_logit(),panel_probit()(FE/RE/CRE),panel_fgls(),interactive_fe()(Bai 2009)panel_unitroot()(IPS/LLC/Fisher/Hadri),mixed()(multilevel/HLM)
New Survival: cox(), kaplan_meier(), survreg(), logrank_test()
New Time Series: var(), granger_causality(), irf(), structural_break(), cusum_test(), engle_granger(), johansen()
New Causal: continuous_did(), rdmc(), rdms() (geographic RD), distributional_te(), mendelian_randomization()
New Design & Data: randomize(), balance_check(), attrition_test(), optimal_design(), mice(), mi_estimate()
New Structural: blp() (BLP demand estimation), frontier() (stochastic frontier)
Smart Workflow Engine (unique to StatsPAI):
recommend()— data + question → estimator recommendation + workflowcompare_estimators()— multi-method comparison with agreement diagnosticsassumption_audit()— one-call assumption testing with remediessensitivity_dashboard()— multi-dimensional sensitivity analysispub_ready()— journal-specific publication readiness checklistreplicate()— built-in famous datasets with replication guides
Interactive Plot Editor: Font presets redesigned to show actual font names; separate font and size presets for independent per-element control.
- OLS
predict(): Out-of-sample prediction viaresult.predict(newdata=) balance_panel(): Keep only units observed in every period- DID/DDD/Event Study weights:
weights=parameter for population-weighted estimation - Matching
ps_poly=: Polynomial propensity score models (Cunningham 2021, Ch. 5) - Synth RMSPE plot:
synthplot(result, type='rmspe')histogram (Abadie et al. 2010) - Graddy (2006) replication: Fulton Fish Market IV example in
sp.replicate() - Numerical validation: Cross-validated against Stata/R reference values
- Theme switching fix: Themes now fully reset rcParams before applying, so switching between themes (e.g. ggplot → academic) correctly updates all visual properties
- Apply button fix: Fixed being clipped on the Layout tab; now pinned to panel bottom
- Error visibility: Widget callback errors now surface in the status bar instead of being silently swallowed
- Auto mode: Always refreshes preview when toggled for immediate feedback
- Theme tab: Moved to first position; color pickers show confirmation feedback
- Code generation: Auto-generate reproducible code with text selection support
Major refactoring and expansion of core modules (+5,800 lines of new code):
- DID: Added Triple Differences (
ddd()), one-calldid_analysis()workflow (auto design detection → Bacon decomposition → estimation → event study → sensitivity), and 8 publication-ready plot functions (parallel_trends_plot,bacon_plot,group_time_plot,enhanced_event_study_plot,treatment_rollout_plot,sensitivity_plot,cohort_event_study_plot) - Synthetic Control: Modular rewrite —
demeaned_synth(),robust_synth()(penalized SCM),gsynth()(Generalized SCM with interactive fixed effects),staggered_synth()(multi-unit staggered adoption),conformal_synth()(distribution-free inference), and comprehensivesynth_plot()/synth_weight_plot()/synth_gap_plot() - Panel: Major expansion of
panel()— Hausman test, Breusch-Pagan LM, Pesaran CD, Wooldridge autocorrelation, panel unit root tests; addedpanel_summary_plot(),fe_plot(),re_comparison_plot() - RD: New
rd_diagnostics()suite — bandwidth sensitivity, placebo cutoffs, donut-hole robustness, covariate balance at cutoff, density test - IV / 2SLS: Rewritten
ivreg()with proper first-stage diagnostics (Cragg-Donald, Kleibergen-Paap), weak IV detection, Sargan-Hansen overidentification test, Anderson canonical correlation test, Stock-Yogo critical values - Matching: Enhanced
match()— added CEM (Coarsened Exact Matching), optimal matching, genetic matching; improved balance diagnostics with Love plot and standardized mean difference - DAG: Expanded
dag()with 15+ built-in example DAGs (dag_example()),dag_simulate()for data generation from causal graphs, backdoor/frontdoor criterion identification - Causal Impact: Enhanced Bayesian structural time-series with automatic model selection and improved inference
- AI Agent Registry: Expanded
list_functions(),describe_function(),function_schema(),search_functions()for LLM/agent tool-use integration - CausalResult: Added
.to_json(),.to_dict(), enhanced.summary()formatting
- Fix PyPI badge displaying stale version
- Meta-Learners: S/T/X/R/DR-Learner for CATE estimation with
compare_metalearners()and CATE diagnostics (gate_test,blp_test) - Neural Causal Models: TARNet, CFRNet, DragonNet for deep CATE estimation
- Causal Discovery:
notears()(continuous DAG optimization),pc_algorithm()(constraint-based) - TMLE: Targeted Maximum Likelihood Estimation with Super Learner
- Policy Learning:
policy_tree()optimal treatment rules,policy_value()evaluation - Conformal Causal: Distribution-free prediction intervals for ITE
- Bayesian Causal Forest:
bcf()with separate prognostic/treatment functions - Dose-Response: Continuous treatment GPS curves
- Bounds: Lee bounds (sample selection), Manski bounds (partial identification)
- Interference:
spillover()direct + indirect effect decomposition - DTR:
g_estimation()multi-stage optimal treatment regimes - Multi-Treatment: AIPW for multi-valued treatments
- Bunching: Kink/notch bunching estimator with elasticity
- Matrix Completion:
mc_panel()nuclear-norm panel estimator - Robustness:
spec_curve(),robustness_report(),subgroup_analysis() - New Regression: DeepIV, Heckman selection, quantile regression, Tobit, Arellano-Bond GMM
- New Diagnostics: E-value, Anderson-Rubin weak IV test, Sensemakr, RD density test
- Other: Entropy balancing, Sun-Abraham event study, Bacon decomposition, HonestDiD
- Post-Estimation:
margins(),marginsplot(),test(),lincom() - Output Tables:
modelsummary(),outreg2(),sumstats(),balance_table(),tab(),coefplot(),binscatter() - Inference:
wild_cluster_bootstrap(),aipw(),ri_test() - New Modules: DML, Causal Forest, Matching (PSM/Mahalanobis), Synthetic Control (ADH + SDID), Panel (FE/RE/FD), Causal Impact, Mediation, Bartik IV
- Diagnostics:
oster_bounds(),mccrary_test(),hausman_test(),het_test(),reset_test(),vif() - Utilities: Variable labeling,
describe(),pwcorr(),winsor(),read_data()
- Core regression:
regress()OLS with robust/clustered/HAC standard errors - Instrumental variables:
ivreg()2SLS - Difference-in-Differences:
did(),did_2x2(),callaway_santanna() - Regression discontinuity:
rdrobust() - Unified
CausalResultobject with.summary(),.plot(),.to_latex(),.to_docx(),.cite()
StatsPAI Inc. is the research infrastructure company behind CoPaper.AI — the AI co-authoring platform for empirical research, born out of Stanford's REAP program.
CoPaper.AI — Upload your data, set your research question, and produce a fully reproducible academic paper with code, tables, and formatted output. Powered by StatsPAI under the hood. copaper.ai

Team:
- Biaoyue Wang — Founder. Economics, Finance, CS & AI. Stanford REAP.
- Dr. Scott Rozelle — Co-founder & Strategic Advisor. Stanford Senior Fellow, author of Invisible China.
-

git clone https://github.com/brycewang-stanford/statspai.git
cd statspai
pip install -e ".[dev,plotting,fixest]"
pytest@software{wang2025statspai,
title={StatsPAI: The Causal Inference & Econometrics Toolkit for Python},
author={Wang, Biaoyue},
year={2026},
url={https://github.com/brycewang-stanford/statspai},
version={1.5.0}
}MIT License. See LICENSE.
GitHub · PyPI · User Guide · CoPaper.AI