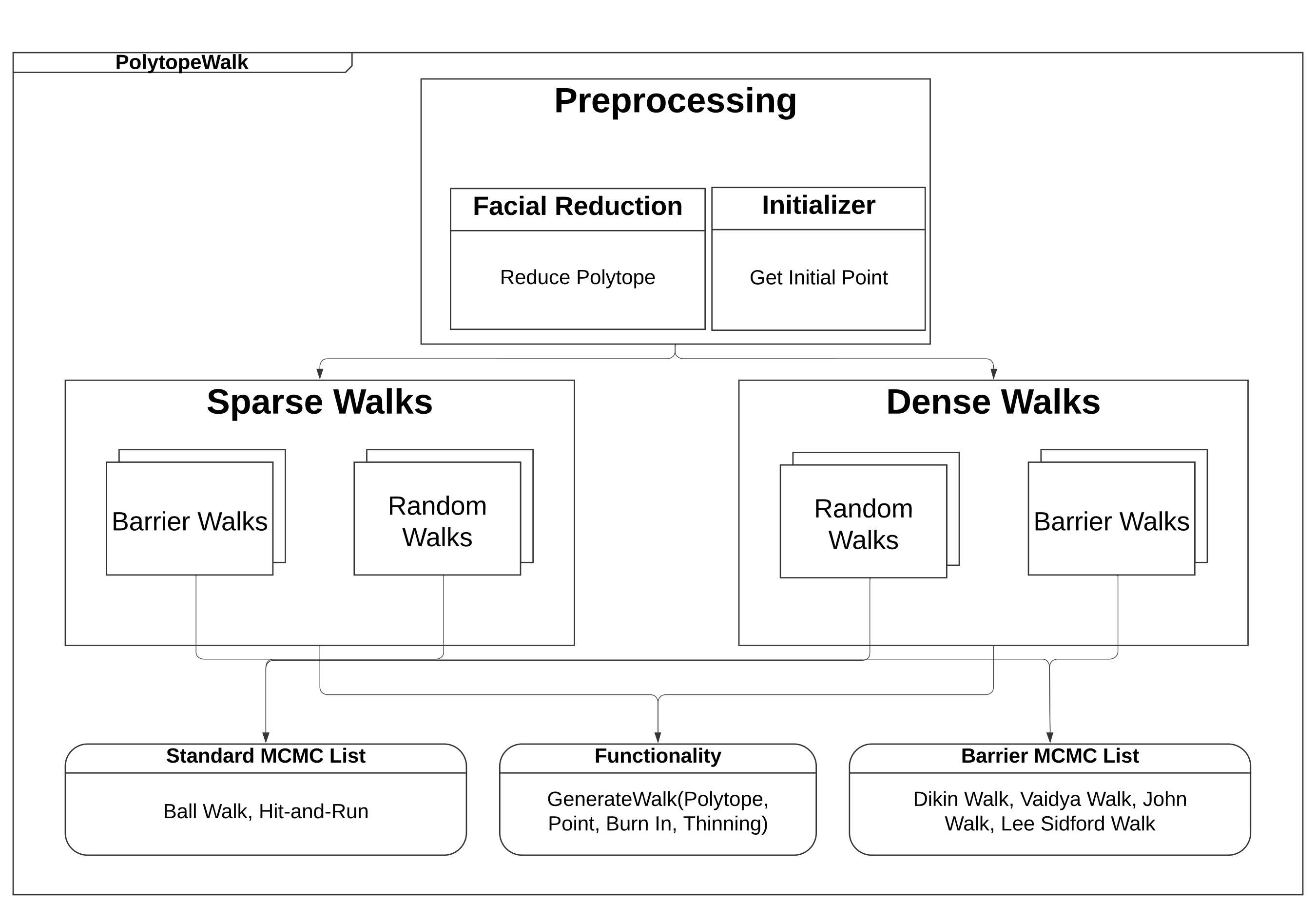
PolytopeWalk is a C++ library for running MCMC sampling algorithms to generate samples from a uniform distribution over a polytope with a Python interface. It handles preprocessing of the polytope (Facial Reduction algorithm) and initialization as well. Current implementations include the Dikin Walk, John Walk, Vaidya Walk, Ball Walk, Lee Sidford Walk, and Hit-and-Run in both the full-dimensional formulation and the sparse constrained formulation. For documentation on all functions/methods, please visit our webpage: https://polytopewalk.readthedocs.io/en/latest/ and read our paper on arXiv here: https://arxiv.org/abs/2412.06629. Finally, for example inputs and outputs, please visit the examples folder, which includes code to uniformly sample from both real-world polytopes from the Netlib dataset and structured polytopes.
Let d be the dimension of the polytope, n be the number of boundaries, and R/r be where the convex body contains a ball of radius r and is mostly contained in a ball of radius R. We implement the following 6 MCMC sampling algorithms for uniform sampling over polytopes.
| Name | Mixing Time | Author |
|---|---|---|
Ball Walk |
Vempala (2005) | |
Hit and Run |
Lovasz (1999) | |
Dikin Walk |
Sachdeva and Vishnoi (2015) | |
Vaidya Walk |
Chen et al. (2018) | |
John Walk |
Chen et al. (2018) | |
Lee Sidford Walk |
Laddha et al. (2019) (conjectured, proof incomplete) |
For each implemented algorithm, we provide the full-dimensional formulation and the sparse constrained formulation. Each polytope can be expressed from 1 formulation to the other. The main benefit of utilizing the constrained formulation is that it maintains sparse operations in A, ensuring scalability in higher dimensions. Many of the netlib dataset sparse polytopes are represented in this formulation. The formulations are specified below.
In the full-dimensional formulation with dense matrix A (
where the polytope is specified with
In the constrained formulation with sparse matrix A (
where the polytope is specified with
In PolytopeWalk, we implement the MCMC algorithms in both the dense, full-dimensional and the sparse, constrained polytope formulation.
PolytopeWalk requires:
- Python (>= 3.9)
- NumPy (>= 1.20)
- SciPy (>= 1.6.0)
If you already have a working installation of NumPy and SciPy, the easiest way to install PolytopeWalk is using pip:
pip install -U polytopewalk- Official source code repo: https://github.com/ethz-randomwalk/polytopewalk
- Download releases: https://pypi.org/project/polytopewalk/
(listed in each of the operating systems)
- macOS:
brew install eigen glpk - Linux:
- Ubuntu
sudo apt-get install -y libeigen3-dev libglpk-dev - CentOS
yum install -y epel-release eigen3-devel glpk-devel
- Ubuntu
- Windows:
choco install eigen -y- Then, install winglpk from sourceforge
git clone https://github.com/ethz-randomwalk/polytopewalk.git
cd polytopewalk
pip install .Only do this, if there is need to run and test C++ code directly. For normal users, we recommend only using the Python interface.
Build with cmake
git clone https://github.com/ethz-randomwalk/polytopewalk.git && cd polytopewalk
cmake -B build -S . & cd build
make
sudo make installThe examples folder provides examples of sampling from both sparse (constrained) and dense (full-dimensional) formulations of the MCMC sampling algorithms as well as testing convergence. We test our random walk algorithms on family of 3 structured polytopes and 3 polytopes from netlib for real-world analysis. The lines below show a quick demonstration of sampling from a polytope using a sparse MCMC algorithm.
import numpy as np
from scipy.sparse import csr_matrix, lil_matrix, csr_array
from polytopewalk.sparse import SparseDikinWalk
def generate_simplex(d):
return np.array([1/d] * d), np.array([[1] * d]), np.array([1]), d, 'simplex'
x, A, b, k, name = generate_simplex(5)
sparse_dikin = SparseDikinWalk(r = 0.9)
dikin_res = sparse_dikin.generateCompleteWalk(10_000, x, A, b, k, burnin = 100, seed = 100)We also demonstrate how to sample from a polytope in a dense, full-dimensional formulation. We additionally introduce the Facial Reduction algorithm, used to simplify the constrained polytope into the full-dimensional form.
import numpy as np
from scipy.sparse import csr_matrix, lil_matrix, csr_array
from polytopewalk.dense import DikinWalk, DenseCenter
from polytopewalk import FacialReduction
def generate_simplex(d):
return np.array([1/d] * d), np.array([[1] * d]), np.array([1]), d, 'simplex'
fr = FacialReduction()
_, A, b, k, name = generate_simplex(5)
dikin = DikinWalk(r = 0.9)
polytope = fr.reduce(A, b, k, sparse = False)
dense_A = polytope.dense_A
dense_b = polytope.dense_b
dc = DenseCenter()
init = dc.getInitialPoint(dense_A, dense_b)
dikin_res = dikin.generateCompleteWalk(1_000, init, dense_A, dense_b, burnin = 100, seed = 100)The tests folder includes comprehensives tests of the Facial Reduction algorithm, Initialization, Weights from MCMC algorithms, and Sparse/Dense Random Walk algorithms in both Python and C++. Our Github package page comes with an automated test suite hooked up to continuous integration after push requests to the main branch.
We provide instructions for locally testing PolytopeWalk in both Python and C++. For both, we must locally clone the repository (assuming we have installed the package already):
git clone https://github.com/ethz-randomwalk/polytopewalk.git
cd polytopewalkIn addition to the requirements from the Developer Installation section, running this code requires a working version of Pandas.
We can run the command:
python -m unittest discover -s tests/python -p "*.py"As mentioned in the Developer Installation section, running this code requires a working version of Eigen and Glpk.
First, we must compile the C++ code :
cmake -B build -S . && cd build
makeThen, we can individually run the test files:
./tests/test_weights
./tests/test_fr
./tests/test_dense_walk
./tests/test_sparse_walk
./tests/test_initFor those wishing to contribute to the software, please feel free to use the pull-request feature on our Github page, alongside a brief description of the improvements to the code. For those who have any issues with our software, please let us know in the issues section of our Github page. Finally, if you have any questions, feel free to contact the authors of this page at this email address: bys7@duke.edu.