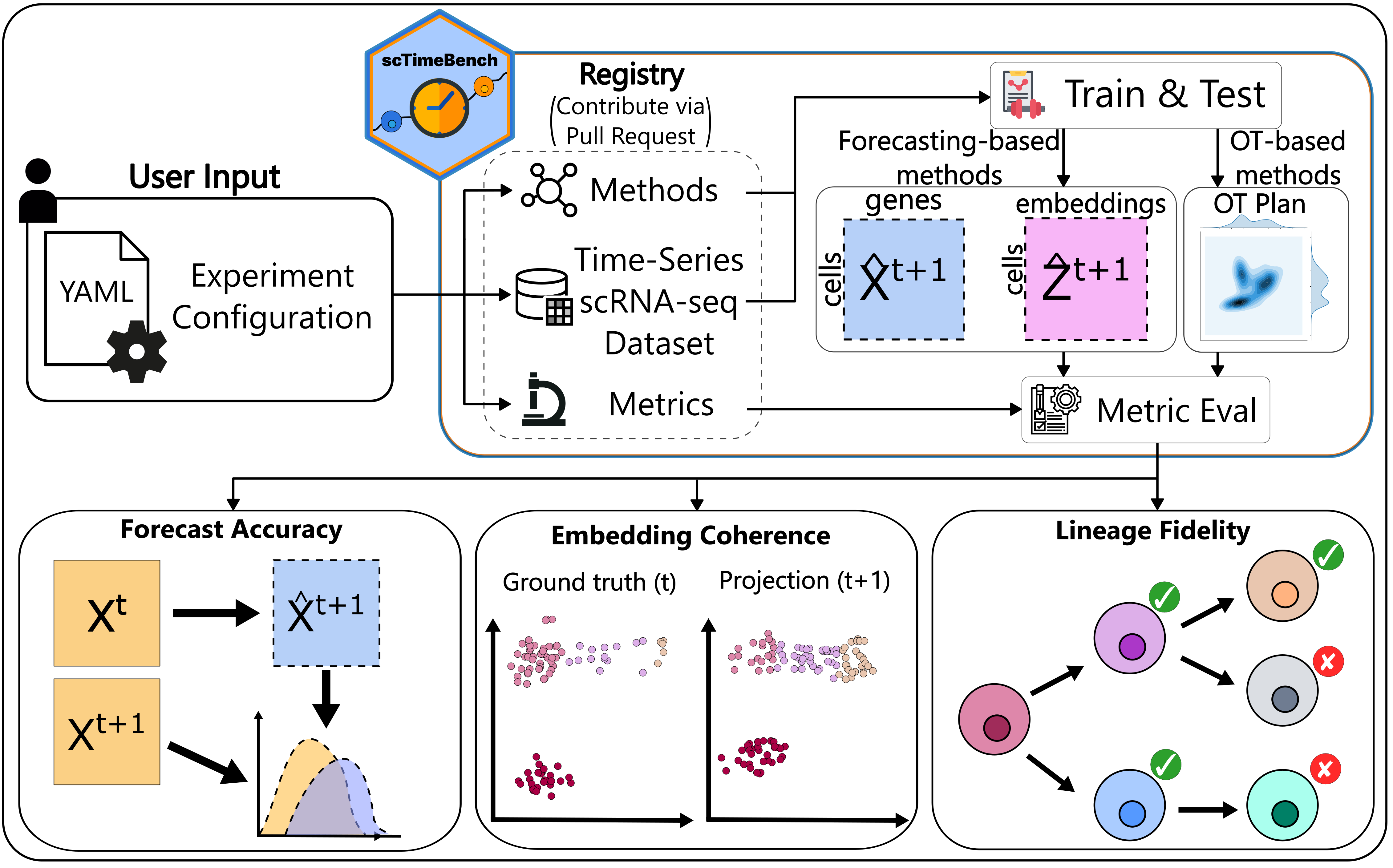
scTimeBench was tested and supported using Python 3.10. If any other version that is 3.10+ does not work when using the benchmark, please submit an issue to this GitHub.
Important Note: this setup is needed twice. Once for the user to run the benchmark metrics, and the second time for the method itself needing to read from scTimeBench.method_utils.method_runner and other important shared constants. In short, this means we need to install scTimeBench into two separate virtual environments:
- Your normal pip installation where you'll be running the benchmark from. This will require the extra "[benchmark]" group installation (pip) or the extra group installation
--extra benchmark(uv). - For each method's virtual environment, you need to install the scTimeBench.
Due to external dependencies and a more complex setup, we have decided to package everything under uv (see: https://github.com/astral-sh/uv). To start with, you need to get all the necessary extern dependencies, which can be done either by running:
git submodule update --init extern/
If you wish to benchmark across all methods, feel free to clone the submodules for all the methods as well with:
git submodule update --init
Then, install uv and run the following:
uv sync --extra benchmark
If you're using uv under a method's virtual environment, either the pip installation or the following will suffice:
uv sync
If the external dependencies such as pypsupertime or sceptic are not used (which they are not used by default), you can install using pip as follows:
pip install scTimeBench[benchmark]
to run the benchmark. For your own method, simply install without the extra benchmarking requirements with
pip install scTimeBench
There are extra dependencies that can be found under pyproject.toml.
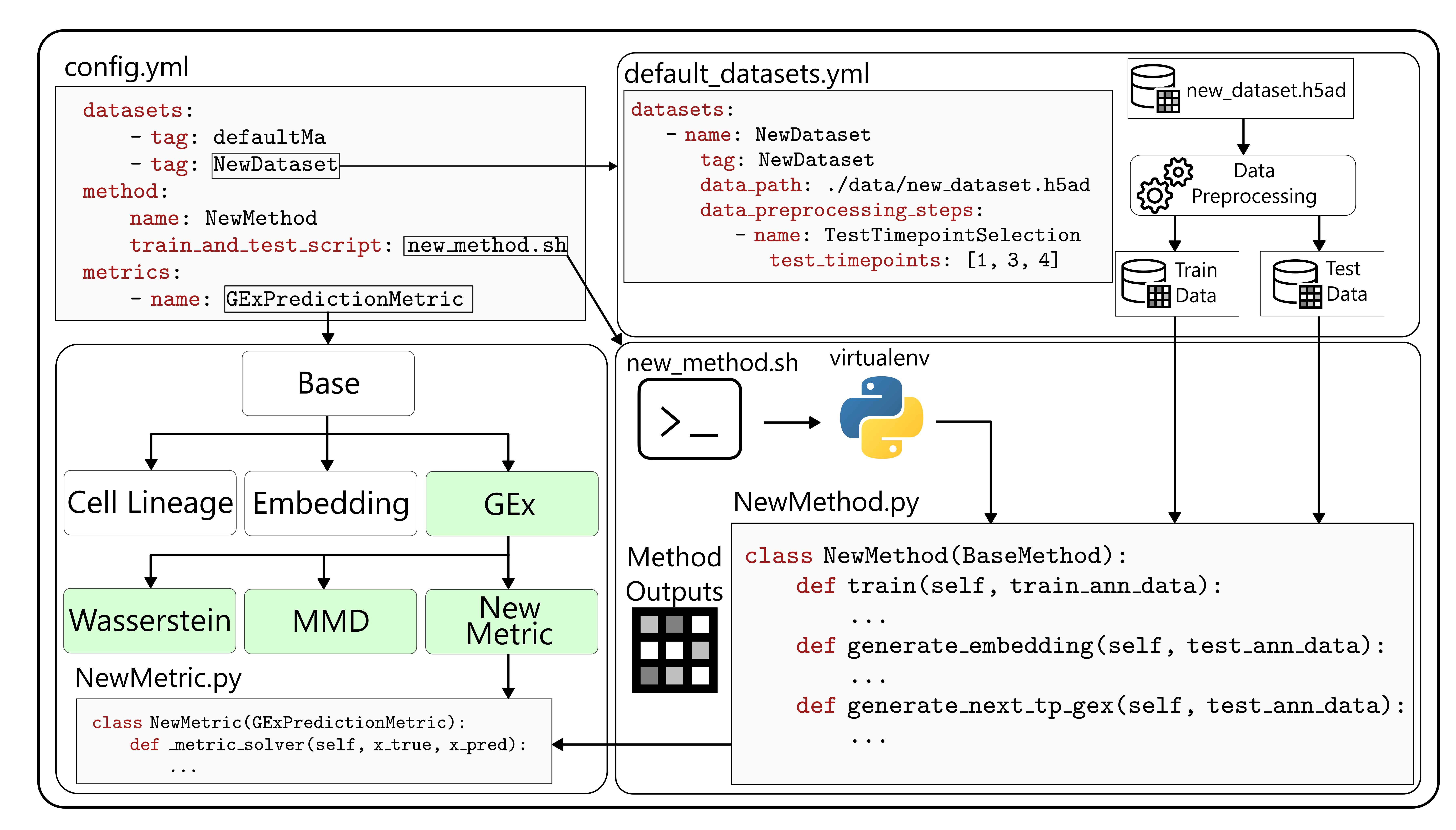 scTimeBench is controlled by a central configuration file which determines which datasets, methods, and metrics to run. An example of this can be found under
scTimeBench is controlled by a central configuration file which determines which datasets, methods, and metrics to run. An example of this can be found under configs/scNODE/gex.yaml.
configs/: possible yaml config files to use as a starting pointextern/: external packages that required edits for compatability such as pyPsupertime and Sceptic.methods/: the different methods that are possible to use, including defined submodules. Add your own methodology here.src/: where the scTimeBench package lies. Seesrc/ReadMe.mdfor more documentation on the modules that exist there.test/: unit tests for each method, dataset, metric, and other important modules.
The entrypoint for the benchmark is defined as scTimeBench. Run scTimeBench --help for more details, or refer to src/scTimeBench/config.py and the documentation.
Run the package with:
scTimeBench --config configs/scNODE/gex.yaml
For a full running example using scNODE, refer to our example Jupyter Notebook.
If you want to contribute, please install the dev environments with:
uv sync --extra dev --extra benchmark
or use pip in editable mode with:
pip install -e ".[dev, benchmark]"
To enable the autoformatting, please run:
pre-commit install
before committing.
Follow our example tutorials on adding new methods, datasets, and metrics in our documentation here: https://li-lab-mcgill.github.io/scTimeBench/.
If your change heavily modifies the architecture, please run the necessary tests under the test/ environment using pytest. Read more on the different available tests under test/ReadMe.md. See more information on the pytest documentation: https://docs.pytest.org/en/stable/. A useful flag is -s to view the entire output of the test.
@article {scTimeBench,
author = {Osakwe, Adrien and Huang, Eric Haoran and Li, Yue},
title = {scTimeBench: A streamlined benchmarking platform for single-cell time-series analysis},
elocation-id = {2026.03.16.712069},
year = {2026},
doi = {10.64898/2026.03.16.712069},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069},
eprint = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069.full.pdf},
journal = {bioRxiv}
}