This repository contains an implementation of the Smith-Waterman algorithm for local sequence alignment. The algorithm is widely used in bioinformatics to find optimal local alignments between two nucleotide or protein sequences.
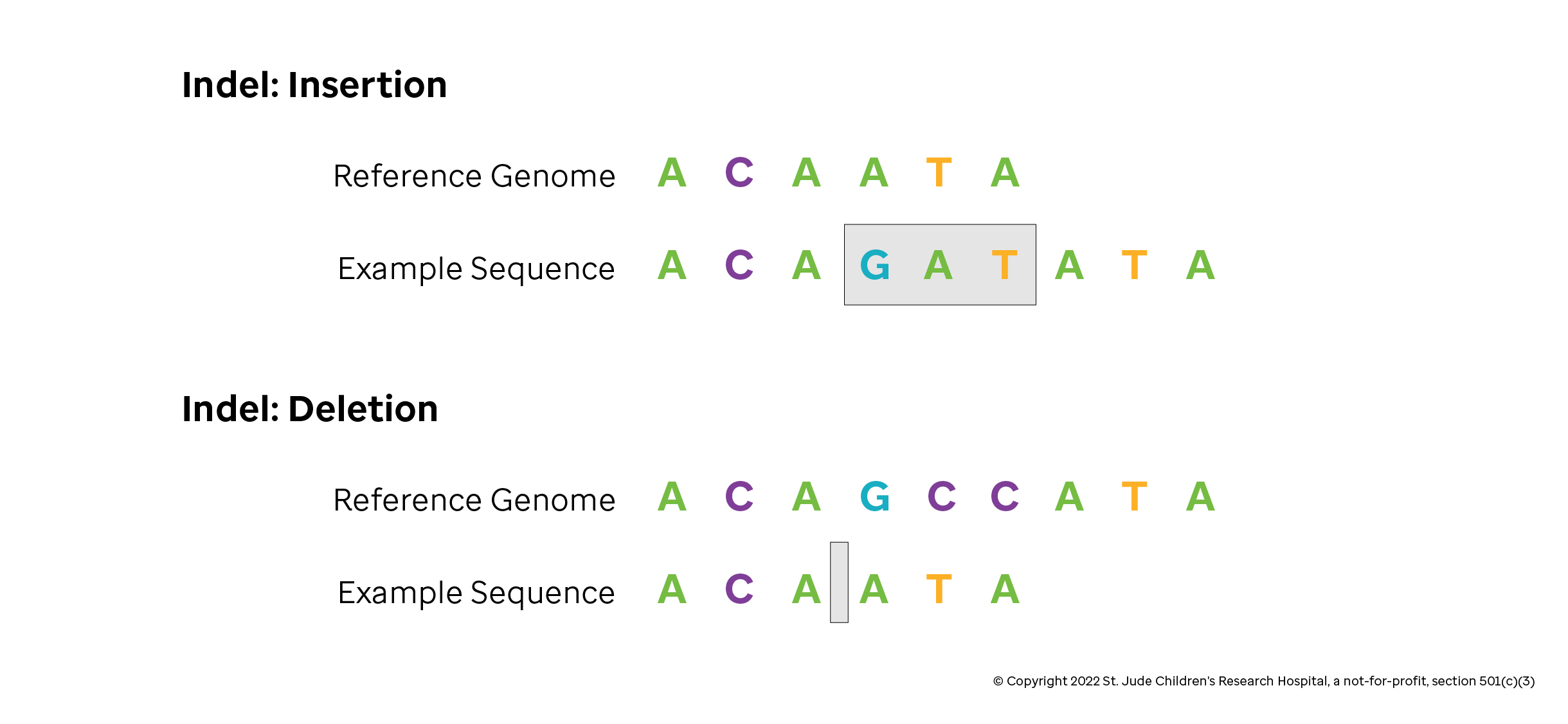
The alignment is performed considering insertions, deletions, and matches/mismatches between two sequences. In biological evolution, mutations can cause insertions or deletions. These are two types of genetic variation in which a specific nucleotide sequence is present (insertion) or absent (deletion).
Hence the algorithm tries to replicate this natural phenomenon introducing a gap event (either an insertion or deletions) to optimize the alignment score when comparing two sequences.
The gap_penalty parameters is critical in Smith-Waterman because it prevents excessive gaps and ensures realistic biological alignments.
- Without a gap penalty → The algorithm might insert too many gaps, artificially maximizing the match score.
- With a gap penalty → The algorithm weighs whether it's better to insert a gap or accept a mismatch.
By fine-tuning its value, you can control how the algorithm balances mismatches vs. gaps for optimal results.
Clone the repository using:
git clone https://github.com/yourusername/smith-waterman.git
cd smith-waterman
python3 -m venv .venv && source .venv/bin/activate
make pip-solveRun the script with sample sequences:
from smith_waterman import water, identity_score
seq1 = "ACACACTA"
seq2 = "AGCACACA"
alignment1, alignment2 = water(seq1, seq2)
print(f"Alignment 1: {alignment1}")
print(f"Alignment 2: {alignment2}")
identity = identity_score(alignment1, alignment2)
print(f"Identity: {identity:.2f}%")Alignment 1: ACACACTA
Alignment 2: AGCACACA
Identity: 87.50%
python -m unittest test_smith_waterman.pyThis project is licensed under the under the Apache License, Version 2.0. See LICENSE for details.
- Smith TF, Waterman MS. Identification of common molecular subsequences. J Mol Biol. 1981 Mar 25;147(1):195-7. doi: 10.1016/0022-2836(81)90087-5. PMID: 7265238.