The first open-source Python package for fully automated CT pelvimetry and body composition analysis.
Automatically measure pelvic dimensions and body composition from CT images — no manual annotation required. Integrates with TotalSegmentator for segmentation and provides a complete DICOM-to-results pipeline.
pip install ctpelvimetry- Fully automated — from DICOM to structured results in one command, no manual landmark placement
- Reproducible — eliminates inter-observer variability inherent in manual CT pelvimetry
- Batch-ready — process hundreds of patients with progress tracking and failure summaries
- Quality-controlled — automatic detection of pelvic rotation, tilt, and sacrum offset with QC figures
- Modular — use the full pipeline or individual analysis functions via CLI or Python API
| Metric | Description |
|---|---|
| ISD (mm) | Inter-Spinous Distance |
| Inlet AP (mm) | Promontory → Upper Symphysis | | Outlet AP (mm) | Coccygeal Apex → Lower Symphysis | | Outlet Transverse (mm) | Intertuberous diameter | | Outlet Area (cm²) | Ellipse approx: π/4 × AP × Transverse | | Sacral Length (mm) | Promontory → Coccygeal Apex | | Sacral Depth (mm) | Max anterior concavity |
| Metric | Description |
|---|---|
| VAT (cm²) | Visceral Adipose Tissue area |
| SAT (cm²) | Subcutaneous Adipose Tissue area |
| V/S ratio | VAT / SAT ratio |
| SMA (cm²) | Skeletal Muscle Area |
Measured at L3 vertebral level and ISD (mid-pelvis) level.
- Per-metric error isolation — failure in one metric does not affect the others
- Quality gates — automatic detection of pelvic rotation, tilt, and sacrum offset
- Batch processing — process hundreds of patients with progress tracking and failure summaries
- QC figures — sagittal combined, extended 3-panel, and body composition overlays
- Modular design — use the full pipeline or individual analysis functions
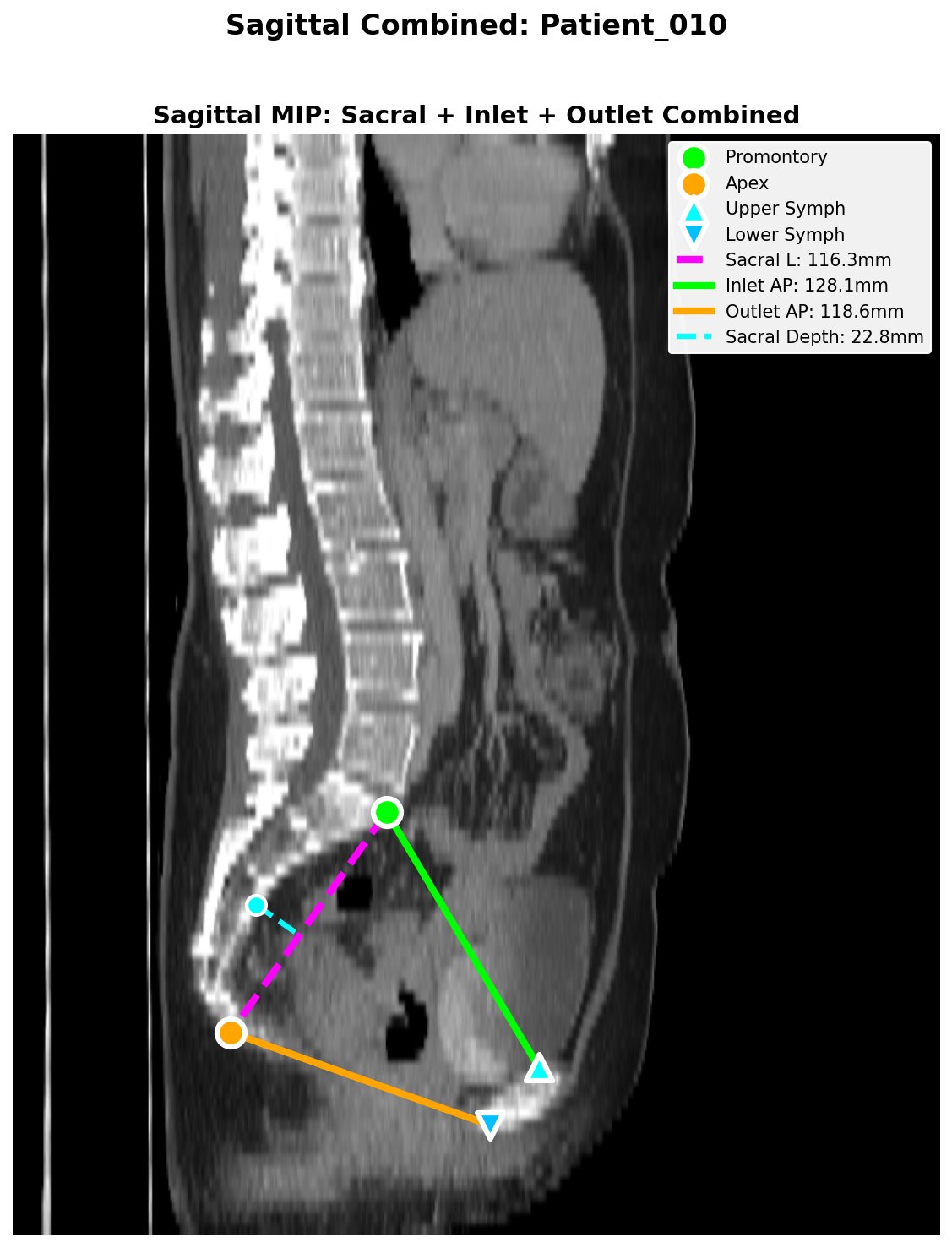
Sagittal QC figure showing automated landmark detection and pelvimetric measurements: sacral length (magenta), inlet AP (green), outlet AP (orange), and sacral depth (cyan).
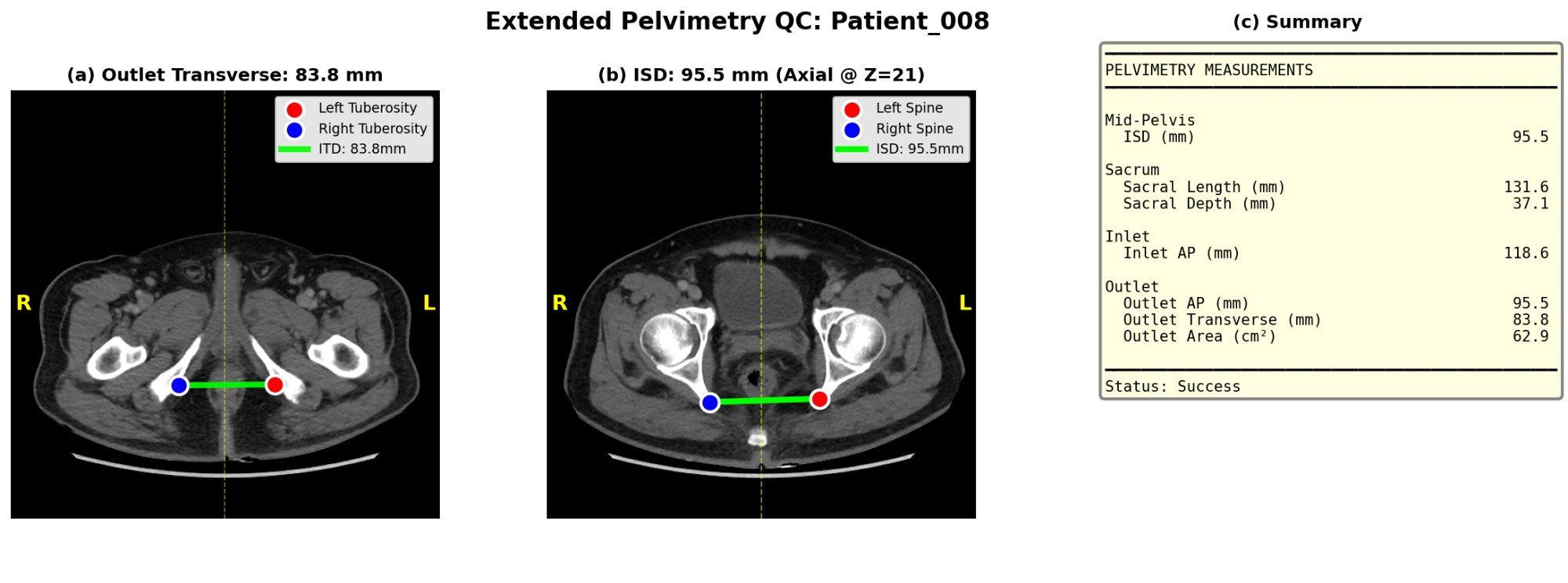
Extended QC panel: (a) outlet transverse diameter at tuberosity level, (b) interspinous distance on axial view, and (c) complete pelvimetry measurement summary.
# Basic install (analyse existing segmentations)
pip install ctpelvimetry
# Full install (includes TotalSegmentator for segmentation)
pip install "ctpelvimetry[seg]"Note: The full install pulls in TotalSegmentator and its PyTorch dependencies. If you only need to analyse pre-existing segmentations, the basic install is sufficient.
| Package | Minimum Version |
|---|---|
| numpy | ≥ 1.24 |
| nibabel | ≥ 5.0 |
| pandas | ≥ 2.0 |
| scipy | ≥ 1.11 |
| matplotlib | ≥ 3.7 |
| tqdm | ≥ 4.60 |
| TotalSegmentator | ≥ 2.0 (optional, pip install ".[seg]") |
ctpelvimetry pelv \
--seg_folder /path/to/segmentations \
--nifti_path /path/to/ct.nii.gz \
--patient Patient_001 \
--output_root ./output --qcctpelvimetry pelv \
--dicom_dir /path/to/Patient_001 \
--output_root ./output \
--patient Patient_001ctpelvimetry body-comp \
--patient Patient_001 \
--seg_root ./batch_output \
--nifti_root ./batch_output \
--pelvimetry_csv ./batch_output/combined_pelvimetry_report.csv \
--output body_comp.csv --qc# Pelvimetry batch
ctpelvimetry pelv \
--dicom_root /path/to/DICOMs \
--output_root ./output \
--start 1 --end 250
# Body composition batch
ctpelvimetry body-comp \
--seg_root ./batch_output \
--nifti_root ./batch_output \
--pelvimetry_csv ./report.csv \
--output body_comp.csv \
--start 1 --end 210 --qc_root ./qcfrom ctpelvimetry import run_combined_pelvimetry, process_single_patient
# Pelvimetry
result = run_combined_pelvimetry(
"Patient_001", "/path/to/seg", "/path/to/ct.nii.gz"
)
# Body composition
result = process_single_patient(
"Patient_001", "/path/to/seg_root",
"/path/to/ct.nii.gz", "/path/to/report.csv"
)ctpelvimetry/
├── __init__.py # Public API
├── config.py # PelvicConfig, constants
├── io.py # Mask loading, coordinate transforms
├── conversion.py # DICOM → NIfTI (dcm2niix)
├── segmentation.py # TotalSegmentator execution
├── landmarks.py # Midline, symphysis, sacral landmarks
├── metrics.py # ISD, ITD, sacral depth
├── body_composition.py # VAT/SAT/SMA analysis
├── qc.py # QC figure generation
├── pipeline.py # run_combined_pelvimetry, run_full_pipeline
├── batch.py # Batch orchestration
└── cli.py # Unified CLI entry point
Contributions are welcome! Please follow these steps:
- Fork the repository
- Create a feature branch (
git checkout -b feature/your-feature) - Commit your changes (
git commit -m "Add your feature") - Push to the branch (
git push origin feature/your-feature) - Open a Pull Request
If you use ctpelvimetry in your research, please cite:
@software{huang2025ctpelvimetry,
author = {Huang, Shih-Feng},
title = {ctpelvimetry: Automated CT Pelvimetry and Body Composition Analysis},
year = {2025},
url = {https://github.com/odafeng/ctpelvimetry},
version = {1.1.0}
}A peer-reviewed manuscript describing ctpelvimetry is currently in preparation. Citation details will be updated upon publication.
This project is licensed under the Apache License 2.0.